PROJECT
Sensing
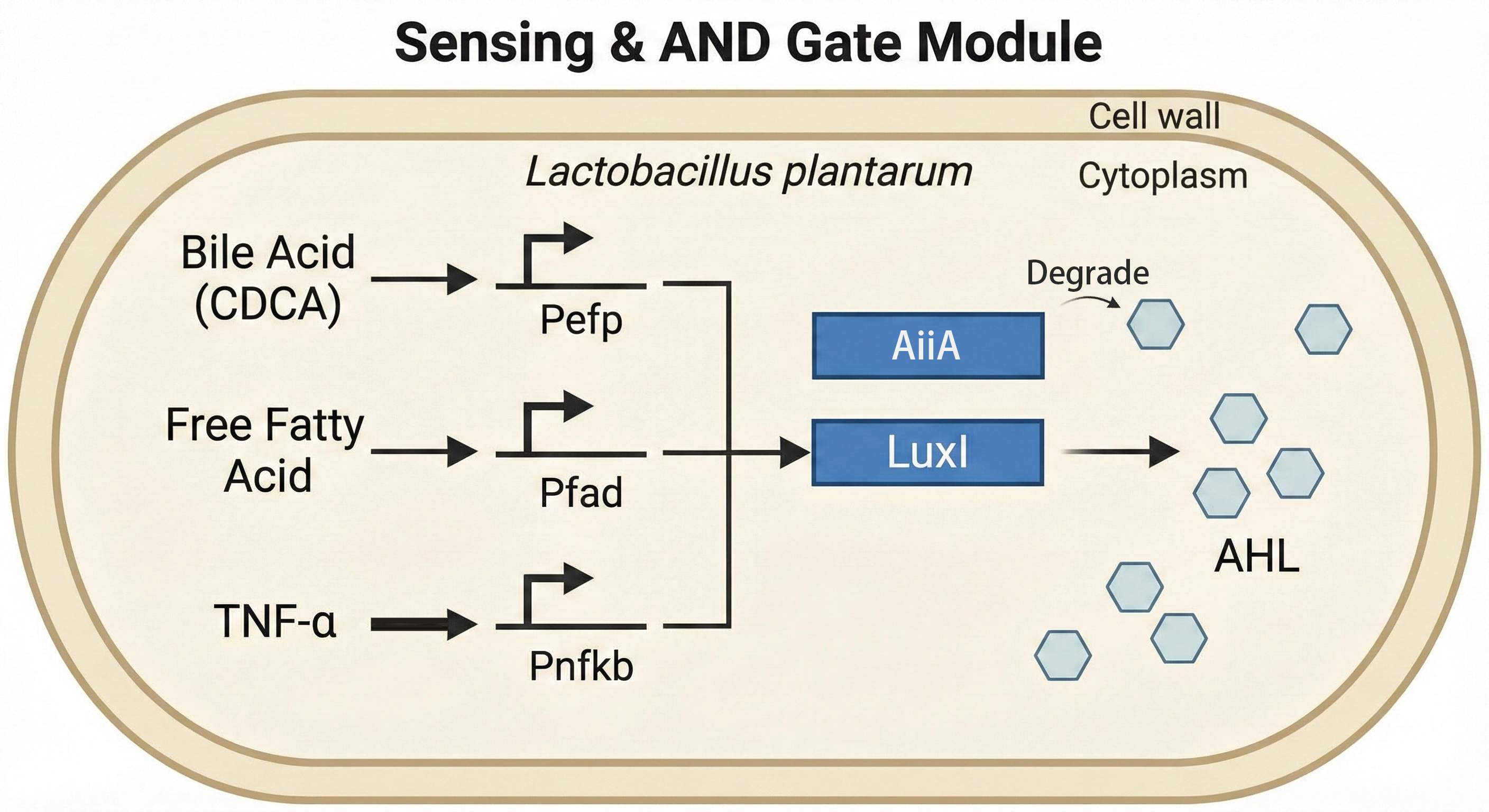
The core innovation of HepaGuard lies in its precise patient‑identification system. We engineered a three‑key AND logic gate that activates therapeutic production only when three NASH‑specific biomarkers simultaneously exceed their thresholds: bile acid (>30 µM via Pefp), free fatty acid (>2 mM via Pfad), and TNF‑α (>50 pg/mL via Pnfkb). These thresholds correspond to bile acid dysmetabolism from liver injury, increased intestinal lipid absorption with barrier dysfunction, and inflammatory markers distinguishing simple steatosis from NASH.
The three promoters drive LuxI expression with weighted contributions: 0.2× (Pefp), 0.5× (Pfad), and 1.0× (Pnfkb). Activation occurs only when the total score exceeds 1.0, requiring either the inflammatory signal Pnfkb together with at least one metabolic signal, or all three simultaneously. This design prevents false activation in healthy individuals—postprandial FFA spikes alone cannot trigger the gate.Pnfkb (TNF-a) must be activated as a master switch, and PEFP (bileacid) and PFAD (FFA-RAct act synergistically as enhancing signals.Three promoters in tandem drive Luxl expression, enabling preciseidentification of Nash patients only. Python simulations based on literature data yield an AUC of 0.947, with 94 % specificity and 89 % sensitivity.
Production
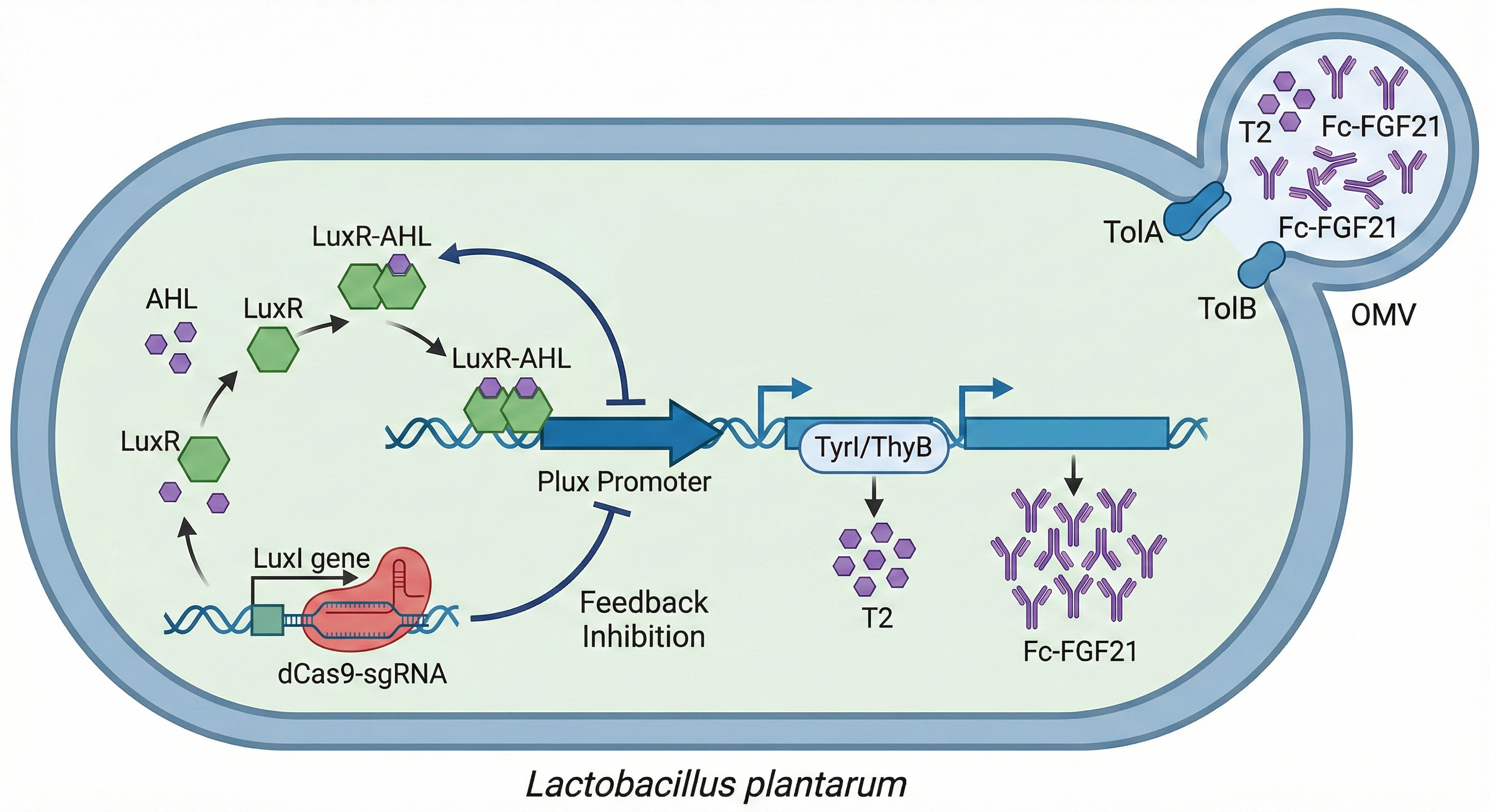
Once activated, the engineered strain initiates a bicistronic operon that simultaneously produces two therapeutic molecules, T2 and Fc‑FGF21, in an optimal 2 : 1 synergistic ratio. T2, a highly selective THR‑β agonist (selectivity >1000‑fold, no cardiac side effects), is synthesised from L‑tyrosine via a two‑step enzymatic pathway using TyrI (a bifunctional iodinase from brown algae) and ThyB (from B. subtilis, responsible for DIT coupling). The final yield reaches 10 mg/L.
Fc‑FGF21 is a fusion protein incorporating a signal peptide for secretion, an Fc fragment to prolong half‑life to 24–48 h, a flexible linker to preserve independent folding, and a His‑tag for easy purification. It improves insulin resistance, suppresses de novo lipogenesis, and protects hepatocytes, with a yield of 5 mg/L. The 2 : 1 ratio is derived from a 2023 Nature Medicine study showing that this combination reduces liver fat by 50 % more than either monotherapy with minimal side effects. Differential RBS strengths and a LuxI/LuxR auto‑calibration system maintain this stable ratio for 48–72 h. Both drugs accumulate in the periplasm and are packaged into outer membrane vesicles (OMVs) induced by TolA/TolB, achieving >60 % encapsulation efficiency, protecting against intestinal proteases, and enabling natural portal‑vein‑to‑liver delivery.
Safety
HepaGuard incorporates quadruple redundant safety locks to ensure automatic clearance after completing its task, preventing long‑term colonisation and ecological risks.
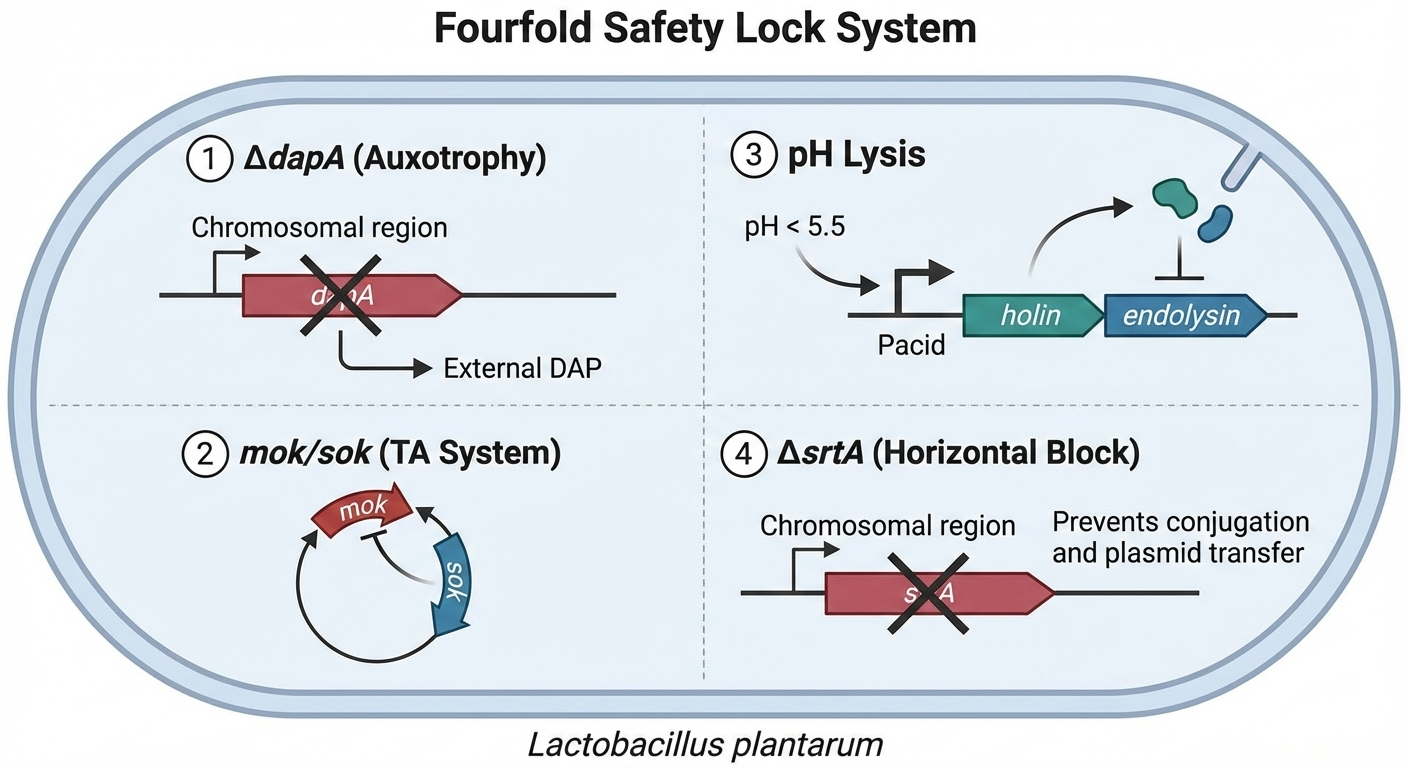
- 🔒 ΔdapA auxotrophy: Genomic deletion of dapA renders the strain dependent on exogenous diaminopimelic acid (DAP). Once the 72‑h DAP supply in the formulation is exhausted, or upon leakage outside the gut, cell‑wall synthesis fails and the bacterium dies.
- 🔒 mok‑sok TA system: Constitutive weak expression of toxin combined with inducible antitoxin RNA leads to toxin accumulation after ~20 divisions due to antitoxin dilution, triggering suicide.
- 🔒 pH‑inducible holin‑endolysin: The Pacid promoter activates at pH < 5.5 (e.g., gastric acid or gut leakage), causing lysis and release of drugs before final self‑destruction.
- 🔒 Horizontal transfer blockade: ΔsrtA knockout reduces surface‑protein anchoring, decreasing conjugation efficiency by 90 %; the pWV01 plasmid has a narrow host range, replicating only in Lactobacillus.
Monte Carlo simulations indicate that while individual lock failure probabilities range from 10⁻³ to 10⁻⁶, the combined failure probability is below 10⁻¹⁸, satisfying FDA standards for live biotherapeutic products. Compared with the chemical drug Resmetirom, HepaGuard offers no cardiac side effects, no hepatic burden, no long‑term residue, and a daily cost of only ¥0.5.
Complete Pathway
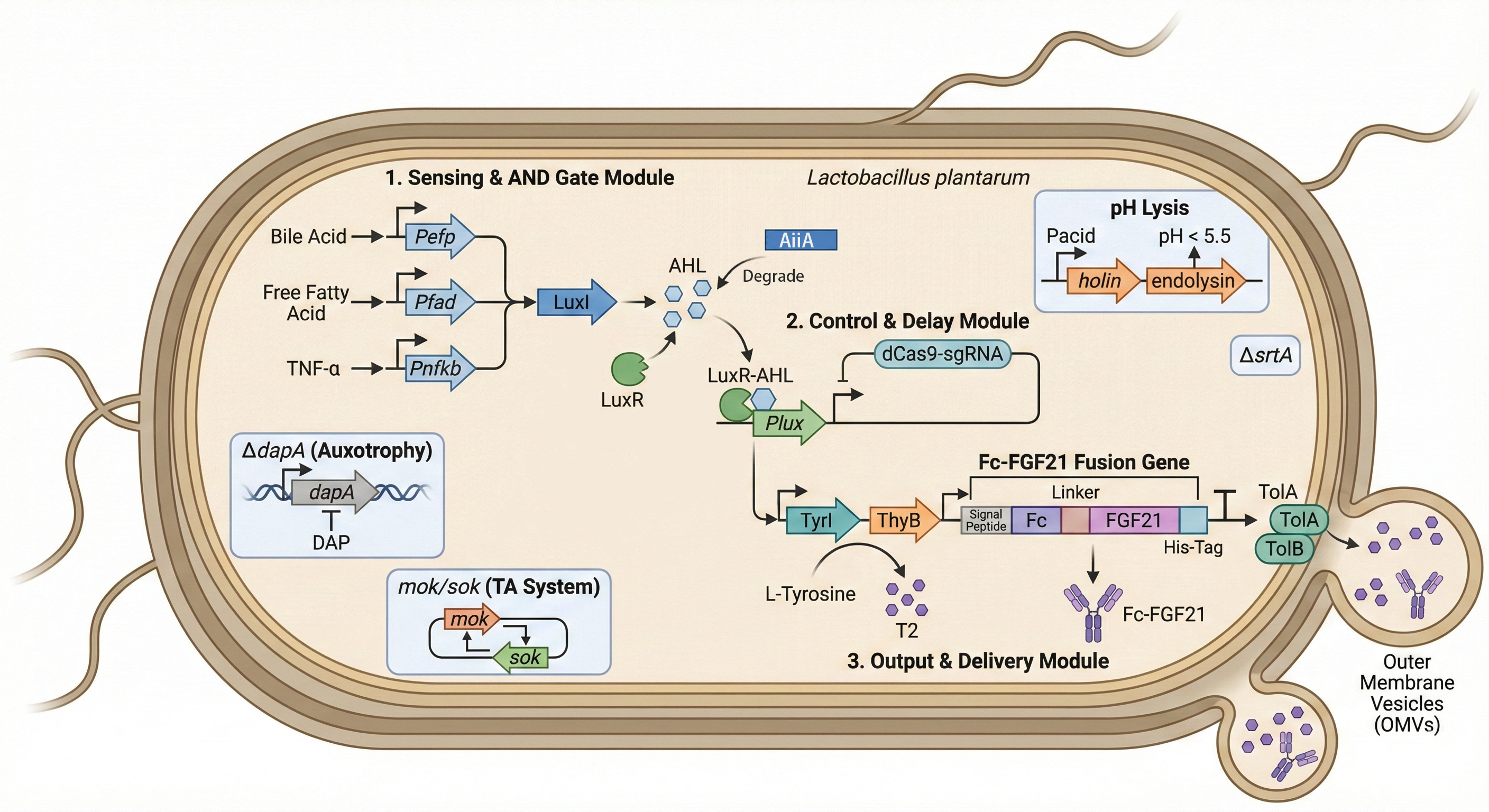
BioBricks
The following BioBrick parts are used in our project:
| Part Name | ID |
|---|---|
| LuxI | BBa_C0061 |
| LuxR | BBa_C0062 |
| Plux | BBa_R0062 |
| Strong RBS | BBa_B0034 |
| Medium RBS | BBa_B0032 |
| Terminator | BBa_B0015 |
| GFP | BBa_E0040 |
| dCas9 | BBa_K1216001 |
References
- Li L, Wang Z, Jiang Y. Global incidence of NAFLD: Sets alarm bells ringing about NAFLD in China again. J Hepatol. 2023;79(4):e164-e165. doi:10.1016/j.jhep.2023.04.030
- Zeng J, Gu C, Wen C, Shen C. The burden of NAFLD (now referred to as MASLD)-related chronic liver disease and cirrhosis from 1990 to 2021 with projections to 2036: a comparative study of global China the United States and India. Lipids Health Dis. 2025;24(1):298. Published 2025 Sep 29. doi:10.1186/s12944-025-02750-z
- Chai SY, Zhang RY, Fernandes G, Zheng YM, Wei L. Epidemiology of metabolic dysfunction-associated steatotic liver disease/metabolic dysfunction-associated steatohepatitis and associated cardiometabolic factors in adults in China (2013-2023): A systematic review and meta-analysis. World J Gastroenterol. 2025;31(46):113608. doi:10.3748/wjg.v31.i46.113608
- Nie Q, Luo X, Wang K, et al. Gut symbionts alleviate MASH through a secondary bile acid biosynthetic pathway. Cell. 2024;187(11):2717-2734.e33. doi:10.1016/j.cell.2024.03.034
- Ding L, Xu JY, Zhang LL, et al. Lactoferrin alleviates non-alcoholic steatohepatitis via remodeling gut microbiota to regulate serotonin-related pathways. J Adv Res. Published online November 16, 2025. doi:10.1016/j.jare.2025.11.034
- Powell EE, Wong VW, Rinella M. Non-alcoholic fatty liver disease. Lancet. 2021;397(10290):2212-2224. doi:10.1016/S0140-6736(20)32511-3
- Noureddin M, Rinella ME, Chalasani NP, et al. Efruxifermin in Compensated Liver Cirrhosis Caused by MASH. N Engl J Med. 2025;392(24):2413-2424. doi:10.1056/NEJMoa2502242
- Moon TS, Lou C, Tamsir A, Stanton BC, Voigt CA. Genetic programs constructed from layered logic gates in single cells. Nature. 2012;491(7423):249-253. doi:10.1038/nature11516
- Miller MB, Bassler BL. Quorum sensing in bacteria. Annu Rev Microbiol. 2001;55:165-199. doi:10.1146/annurev.micro.55.1.165
- Feng H, Guo J, Wang T, Zhang C, Xing XH. Guide-target mismatch effects on dCas9-sgRNA binding activity in living bacterial cells. Nucleic Acids Res. 2021;49(3):1263-1277. doi:10.1093/nar/gkaa1295
- Zhao Y, Li L, Zheng G, et al. CRISPR/dCas9-Mediated Multiplex Gene Repression in Streptomyces. Biotechnol J. 2018;13(9):e1800121. doi:10.1002/biot.201800121
- Byun G, Yang J, Seo SW. CRISPRi-mediated tunable control of gene expression level with engineered single-guide RNA in Escherichia coli. Nucleic Acids Res. 2023;51(9):4650-4659. doi:10.1093/nar/gkad234
- Reimer SL, Beniac DR, Hiebert SL, et al. Comparative Analysis of Outer Membrane Vesicle Isolation Methods With an Escherichia coli tolA Mutant Reveals a Hypervesiculating Phenotype With Outer-Inner Membrane Vesicle Content. Front Microbiol. 2021;12:628801. Published 2021 Mar 5. doi:10.3389/fmicb.2021.628801
- De Filippis F, Pasolli E, Ercolini D. The food-gut axis: lactic acid bacteria and their link to food, the gut microbiome and human health. FEMS Microbiol Rev. 2020;44(4):454-489. doi:10.1093/femsre/fuaa015
- Li M, Mao B, Tang X, et al. Lactic acid bacteria derived extracellular vesicles: emerging bioactive nanoparticles in modulating host health. Gut Microbes. 2024;16(1):2427311. doi:10.1080/19490976.2024.2427311
- Song J, Zhai P, Zhang Y, et al. The Aspergillus fumigatus Damage Resistance Protein Family Coordinately Regulates Ergosterol Biosynthesis and Azole Susceptibility. mBio. 2016;7(1):e01919-15. Published 2016 Feb 23. doi:10.1128/mBio.01919-15
- Le Rhun A, Tourasse NJ, Bonabal S, Iost I, Boissier F, Darfeuille F. Profiling the intragenic toxicity determinants of toxin-antitoxin systems: revisiting hok/Sok regulation. Nucleic Acids Res. 2023;51(1):e4. doi:10.1093/nar/gkac940
- Peng J, Triplett LR, Schachterle JK, Sundin GW. Chromosomally Encoded hok-sok Toxin-Antitoxin System in the Fire Blight Pathogen Erwinia amylovora: Identification and Functional Characterization. Appl Environ Microbiol. 2019;85(15):e00724-19. Published 2019 Jul 18. doi:10.1128/AEM.00724-19
- Zhang M, Wang Y, Chen J, et al. Identification and Characterization of a New Type of Holin-Endolysin Lysis Cassette in Acidovorax oryzae Phage AP1. Viruses. 2022;14(2):167. Published 2022 Jan 18. doi:10.3390/v14020167
- Bierne H, Mazmanian SK, Trost M, et al. Inactivation of the srtA gene in Listeria monocytogenes inhibits anchoring of surface proteins and affects virulence. Mol Microbiol. 2002;43(4):869-881. doi:10.1046/j.1365-2958.2002.02798.x