WET LAB

Plasmid Construction and Verification
Objective: Assemble the complete HepaGuard genetic circuit: pSIP411‑Pefp‑Pfad‑Pnfkb‑Plux‑TyrI‑ThyB‑FcFGF21, integrating three sensing promoters, AND‑gate controlled LuxI, dual‑drug operon (TyrI‑ThyB for T2, Fc‑FGF21), and safety modules.
Materials:
- Backbone: pSIP411 (chloramphenicol resistance, suitable for Lactobacillus).
- Gene fragments: TyrI (synthesized from Laminaria digitata), ThyB (from B. subtilis), Fc‑FGF21 (gene synthesis with signal peptide, Fc domain, linker, His‑tag).
- Promoters: Pefp, Pfad, Pnfkb (iGEM Registry or synthesis).
- Enzymes: BsaI (Golden Gate), T4 DNA ligase.
- Competent cells: E. coli DH5α (cloning), L. plantarum LP‑115 (expression).
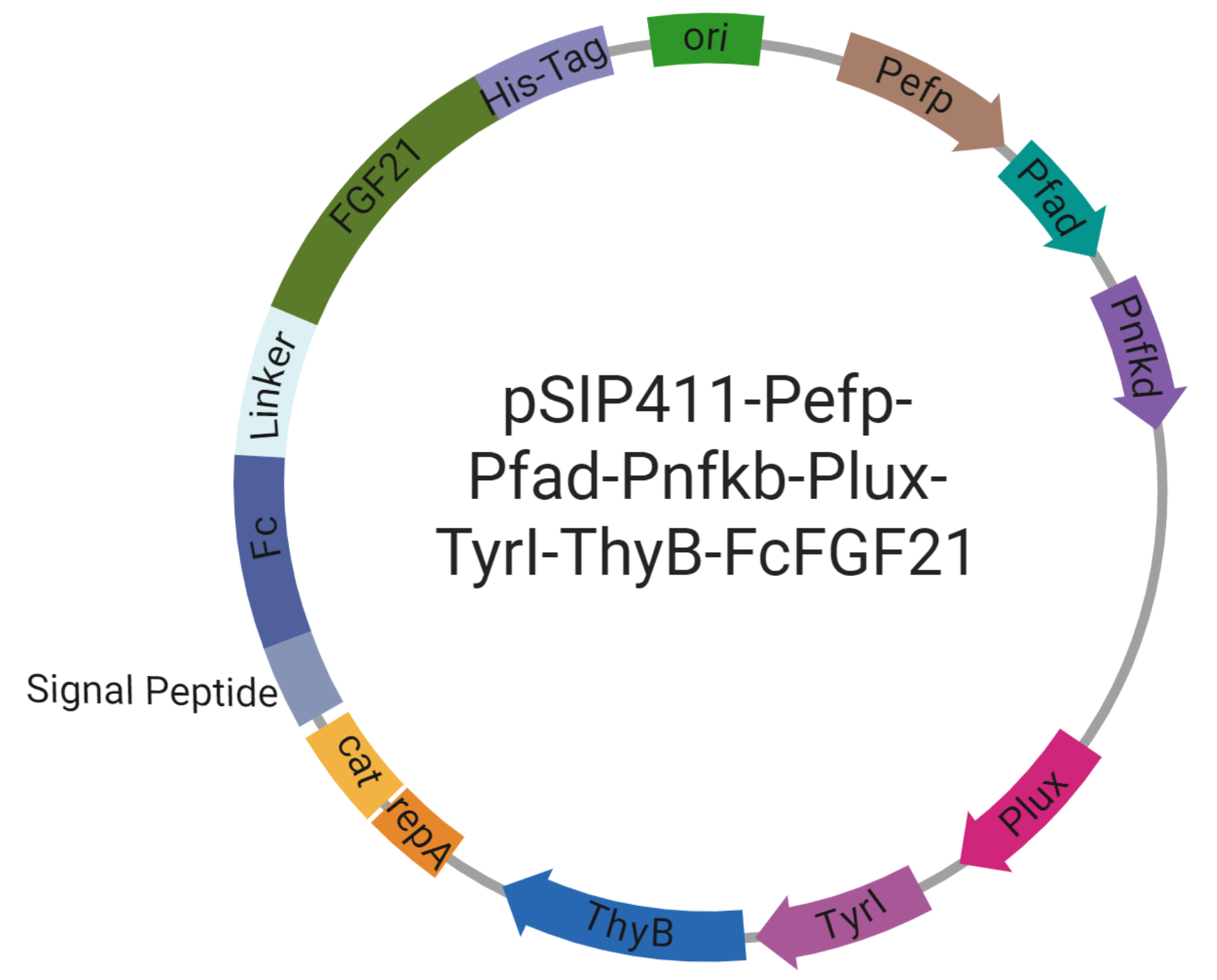
Expected Results:
- Positive clone rate >50% (colony PCR).
- Sequencing accuracy >90% (no mutations in promoters/coding frames).
- Transformation efficiency in L. plantarum >10⁴ CFU/μg.
Validation: Colony PCR + sequencing report.
AND Gate Logic Verification
Objective: Confirm the three‑key AND gate activates only when multiple NASH signals exceed threshold, preventing false activation by single signals.
Materials:
- Reporter strain: LP‑115/pSIP411‑AND‑gfp (complete three‑input circuit).
- Inducer combinations: CDCA (0/50 μM), oleic acid (0/5 mM), TNF‑α (0/100 pg/mL).
- Detection: flow cytometry (single‑cell) or microplate reader (bulk).
Expected Results:
- Triple induction → fluorescence >20‑fold over background.
- Single inducers → <2‑fold increase.
- Pairwise combinations → intermediate responses depending on promoter weights.
- Hill coefficient >2, digital‑like threshold.
- Truth table matches computational predictions (Pnfkb=1.0, Pfad=0.5, Pefp=0.2, total >1.0 activates).
Validation: Fluorescence bar graph for 8 combinations, truth table, dose‑response curves.
T2 and FGF21 Production Quantification
Objective: Quantify T2 and Fc‑FGF21 production in activated HepaGuard, verifying optimal 2:1 synergistic ratio.
Materials:
- Engineered strain: LP‑115/pSIP411‑complete circuit.
- Induction: CDCA 50 μM + oleic acid 5 mM + TNF‑α 100 pg/mL in MRS + 5 mM NaI (for T2 synthesis).
- Instrumentation: LC‑MS/MS (triple quadrupole) for T2, His‑tag capture ELISA for FGF21.
- Time points: 12, 24, 48, 72 h.
Expected Results:
- Total T2 yield: 8–12 mg/L (peak at 48 h).
- FGF21 yield: 4–6 mg/L.
- Stable T2:FGF21 ratio ≈ 1.5–2.5:1 (matching 2:1 synergy).
- Combined therapeutic protein >10 mg/L, maintained 48–72 h.
Validation: Standard curves, chromatograms, ELISA calibration, ratio time‑course.
OMV Isolation and Characterization
Objective: Verify that OMVs encapsulate therapeutic cargo, protect from degradation, and target liver naturally.
Materials:
- Engineered bacteria induced 24 h.
- Differential centrifugation: 10,000×g (remove cells) → 100,000×g (pellet OMVs).
- Characterization: TEM (morphology), DLS (size, PDI), Western blot (OmpA marker).
- Encapsulation efficiency: drug concentration in purified OMVs vs. total production.
Expected Results:
- Spherical vesicles, 50–100 nm diameter (TEM).
- Narrow size distribution (PDI <0.3).
- OmpA signal confirmed (Western blot).
- Encapsulation: T2 >60%, FGF21 >50%.
- Yield >10¹² particles/mL.
Validation: TEM images, DLS histograms, Western blot bands, encapsulation efficiency data.
In Vitro Hepatocyte Uptake Verification
Objective: Demonstrate OMV internalization by hepatocytes in a NASH‑mimetic model.
Materials:
- HepG2 cells cultured in DMEM + 10% FBS.
- NASH model: 0.5 mM palmitic acid‑BSA for 48 h.
- OMVs labeled with DiO or PKH67 (10–100 μg/mL, 1–4 h).
- Detection: flow cytometry (% DiO⁺, MFI) and confocal microscopy.
Expected Results:
- Dose‑dependent uptake: >40% positive cells at 10 μg/mL, 4 h.
- Rapid internalization within 1 h, plateau at 4 h.
- Punctate cytoplasmic fluorescence (endosomes), not surface binding.
- Enhanced uptake in steatotic cells.
- No cytotoxicity (LDH assay).
Validation: Flow scatter plots, confocal images, LDH release data.
3D Hepatospheres NASH Model
Objective: Validate therapeutic efficacy of OMV‑encapsulated drugs in a physiologically relevant 3D liver model.
Materials:
- HepG2 spheroids formed in ultra‑low attachment plates (72 h, 200–300 μm diameter).
- NASH induction: 1 mM FFA (oleate:palmitate 2:1) + 50 ng/mL TNF‑α for 48 h.
- Treatment groups: control, NASH, NASH + empty OMVs, NASH + T2‑OMVs, NASH + FGF21‑OMVs, NASH + dual‑drug OMVs (2:1).
- Endpoints: Oil Red O (lipid quantification), ELISA (TNF‑α, IL‑6), live/dead staining.
Expected Results:
- NASH spheroids show >3‑fold lipid accumulation vs. control.
- Dual‑drug OMVs reduce lipid area >40% compared to NASH.
- Inflammatory cytokines decrease >50%.
- Viability >80% in all groups.
- Single‑drug treatments show intermediate efficacy (synergy confirmed).
Validation: Oil Red O quantification, cytokine ELISA, viability staining.
Dual Safety Lock Verification
Objective: Confirm that redundant safety systems prevent horizontal gene transfer and maintain genetic stability over therapeutic duration.
Materials:
- Horizontal transfer: co‑culture HepaGuard (donor, Cmᴿ) with E. coli MG1655 (recipient, Cmˢ) on solid medium 24 h; select transconjugants on Cm plates, confirm by PCR.
- Stability: continuous passage in MRS + DAP for 20 generations (72 h); sample every 4 generations; re‑induce AND gate; sequence key genes (LuxI, TyrI, FGF21, safety modules).
Expected Results:
- Transfer frequency <10⁻⁶ per donor (ΔsrtA + narrow‑host‑range pWV01 effective).
- After 20 generations, AND gate induction efficiency decrease <5%.
- No mutations in critical regions (sequencing).
- Plasmid retention >95% (mok/sok TA system).
Validation: Conjugation plates, PCR, sequencing data, antibiotic sensitivity tests.