Genetic Circuit & Parts
1. System Overview
We designed a dual-target "AND" gate detection system based on RNA aptamers for the simultaneous detection of NFL (neurofilament light chain) and GFAP (glial fibrillary acidic protein) in blood. The system consists of two plasmids:
- Plasmid 1 (pUC19 backbone): Expresses the blocking strand RNA1 (containing a T7 promoter complementary sequence, GFAP aptamer, and NFL aptamer).
- Plasmid 2 (pUC57 backbone): Reporter plasmid containing a T7 promoter and the sgRNA gene.
When NFL and GFAP are both present, they bind to their respective aptamers on RNA1, causing RNA1 to dissociate from the T7 promoter on plasmid 2. T7 RNA polymerase then initiates sgRNA transcription. The sgRNA activates Cas13, which cleaves the reporter probe, ultimately producing a visible readout on a lateral flow test strip.
2. Mechanism
OFF State (No Target or Single Target Only)
Under arabinose induction, plasmid 1 transcribes RNA1 (containing the blocker sequence and dual aptamers). Through its T7 complementary sequence, RNA1 forms a stable duplex with the T7 promoter DNA on plasmid 2 (ΔG = -27.35 kcal/mol, RNAcofold modeling result), physically blocking T7 RNA polymerase binding. Transcription is terminated, and no sgRNA is produced. Cas13 remains inactive, and the reporter probe remains intact.
ON State (Both NFL and GFAP Present)
When both NFL and GFAP are present in the sample:
- NFL binds to the NFL aptamer on RNA1.
- GFAP binds to the GFAP aptamer on RNA1.
- Protein binding induces a conformational change in RNA1, causing it to dissociate from the T7 promoter DNA.
- T7 RNA polymerase binds to the promoter and transcribes sgRNA.
- sgRNA activates Cas13, which cleaves the reporter probe.
Test Strip Interpretation
| State | sgRNA | Cas13 Activity | Probe Status | T Line | C Line | Interpretation |
|---|---|---|---|---|---|---|
| No target / Single target | Absent | Inactive | Intact | Visible | Visible | Negative (no dual target) |
| Dual target present | Present | Active | Cleaved | Invisible | Visible | Positive (NFL+GFAP both present) |
Note: The C line serves as a control and should be visible in both positive and negative results; its absence indicates an invalid test strip.
3. Parts List
| Part | Type | Source/Biobrick | Function | Reference |
|---|---|---|---|---|
| T7 promoter | Promoter | BBa_K5498011 | Drives transcription of RNA1 and sgRNA | iGEM Registry |
| T7 terminator | Terminator | BBa_25ECYPMT | Terminates transcription | iGEM Registry |
| pBAD promoter | Promoter | BBa_25T3HMKA | Arabinose-inducible expression of RNA1 | iGEM Registry |
| NFL aptamer (MN734) | Aptamer | To be submitted | Specifically binds NFL protein, Kd = 8.1 nM | Matsumoto et al., 2026 |
| GFAP aptamer | Aptamer | To be submitted | Specifically binds GFAP protein, Kd = 0.62 μM | Cao et al., 2025 |
| Blocker sequence | Blocking sequence | To be submitted | Complementary to T7 promoter, mediates RNA1 binding | Team design |
| sgRNA | Reporter gene | To be submitted | Activates Cas13, triggers signal | Team design |
| Cas13 | Effector protein | Literature source | Cleaves RNA probe upon sgRNA activation | Literature |
4. Plasmid Design
| Plasmid | Backbone | Resistance | Origin of Replication | Insert | Function |
|---|---|---|---|---|---|
| Plasmid 1 | pUC19 | Ampicillin | pMB1 | pBAD promoter - blocker - (G?S)? linker - GFAP aptamer - (G?S)? linker - NFL aptamer - T7 terminator | Arabinose-inducible expression of RNA1 |
| Plasmid 2 | pUC57 | Kanamycin | pUC | T7 promoter - sgRNA - T7 terminator | Reporter plasmid, regulated by RNA1 |
Plasmid Maps
Plasmid 1 (pUC19):
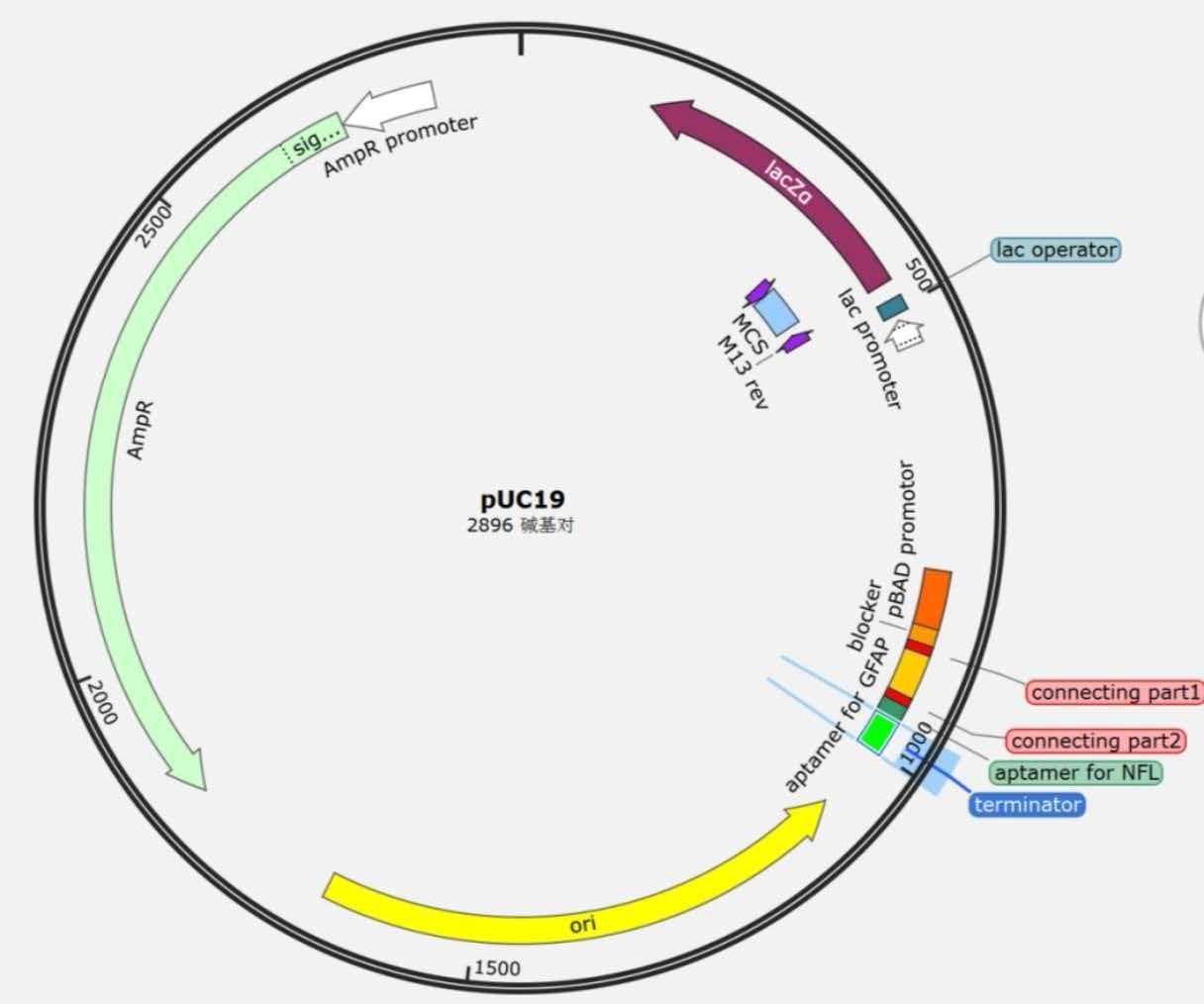
Plasmid 2 (pUC57):
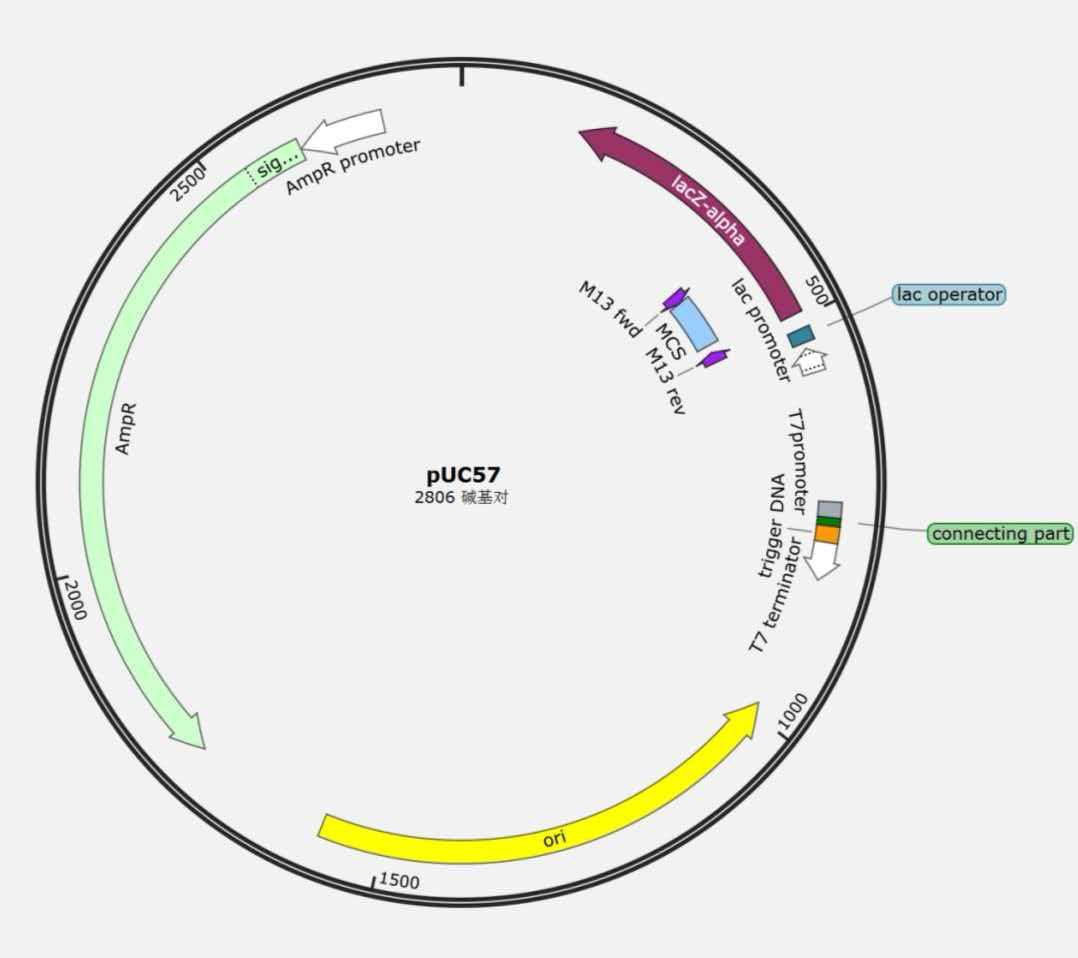
RNA1 Sequence: The full-length DNA sequence has been designed and is available in the Wiki appendix. It contains the T7 promoter complementary region, both aptamers, and linker sequences.
5. Workflow
- Sample Processing: Mix blood sample with detection reagents (containing Cas13, reporter probe, etc.), incubate at 37°C for 30 minutes.
- Apply to Test Strip: Apply the reaction mixture to the sample pad of a lateral flow test strip.
- Result Readout: Visually inspect after 5–15 minutes.
- Positive (NFL+GFAP both present): C line visible, T line invisible.
- Negative (no target or single target): Both C line and T line visible.
- Invalid: C line invisible.
6. Key Supporting Data
| Data | Method | Result | Source |
|---|---|---|---|
| OFF-state binding energy | RNAcofold | ΔG = -27.35 kcal/mol | Modeling |
| NFL aptamer affinity | Literature | Kd = 8.1 nM (MN734) | Matsumoto et al., 2026 |
| GFAP aptamer affinity | Literature | Kd = 0.62 μM | Cao et al., 2025 |
| NFL aptamer–protein binding | HDOCK | -288.23 | Modeling |
| GFAP aptamer–protein binding | HDOCK | -301.91 | Modeling |
| "AND" gate validation | ODE modeling | sgRNA produced only with both targets | Modeling |
| Detection performance | ODE modeling | LOD = 0.57 pM, linear range 1–10 nM | Modeling |
7. References
- Kim KY, Shin KY, Chang KA. GFAP as a Potential Biomarker for Alzheimer's Disease: A Systematic Review and Meta-Analysis. Cells. 2023;12(9):1309. (GFAP biomarker rationale)
- Wang X, Shi Z, Qiu Y, Sun D, Zhou H. Peripheral GFAP and NfL as early biomarkers for dementia: longitudinal insights from the UK Biobank. BMC Med. 2024;22(1):192. (NFL+GFAP combined detection rationale)
- Matsumoto M, Ikebukuro K, Tsukakoshi K. Competitive-SELEX discovery of DNA aptamers selective for neurofilament light chain in human plasma. Biochem Biophys Res Commun. 2026;796:153151. (NFL aptamer source)
- Buonocore J, et al. Plasma NfL and GFAP in the preclinical stages of neurodegenerative diseases: insights from the UK Biobank. J Neurol. 2025;272(12):755. (UK Biobank study)
- Cao Y, Lin Y, Chen L, et al. Screening of glial fibrillary acidic protein specific aptamer and application in the development of fluorescent biosensor based on isothermal amplification strategy. Chin J Anal Chem. 2025. doi:10.1016/j.cjac.2025.100184. (GFAP aptamer source)
8. Additional Notes
- Biobrick Submission Plan: The novel aptamer parts (NFL aptamer, GFAP aptamer) and blocker sequence designed in this project will be submitted to the iGEM Registry after the competition.
- Ethical Compliance: This system is designed for in vitro detection only, involving no animal or human experiments, in compliance with iGEM ethical guidelines.
- Raw Data: All modeling data and sequence information have been archived and are available in the corresponding Wiki pages.