Project
Project Introduction
Alzheimer's disease (AD) affects over 10 million people in China. However, most patients are diagnosed only at moderate or advanced stages, mainly due to significant limitations of current diagnostic methods: invasive cerebrospinal fluid (CSF) puncture, costly PET-CT, and subjective cognitive scale assessment.
AlzBiome is a dual-target in vitro diagnostic system designed to simultaneously detect two blood-based biomarkers: NFL (neurofilament light chain, reflecting axonal injury) and GFAP (glial fibrillary acidic protein, reflecting neuroinflammation). Using an RNA aptamer-based "AND" gate design, the system is intended to generate a detectable signal only when both targets are present, thereby reducing the risk of false positives compared with single-target detection.
Current stage: Complete design and computational validation. The full-length DNA sequence has been designed and is ready for experimental validation pending laboratory access.
Project Design
1. Core Design Principle
We designed a competitive transcription regulation system based on RNA aptamers to achieve dual-target "AND" gate synergistic detection of NFL and GFAP.
DNA template design (5'→3'):
[T7 promoter] — [Blocker] — [10T linker] — [GFAP aptamer] — [10T linker] — [NFL aptamer]
The blocker sequence is complementary to the T7 promoter region, enabling RNA1 to bind and block transcription. The two aptamers are connected by flexible linkers to allow independent protein binding.
2. Plasmid Design
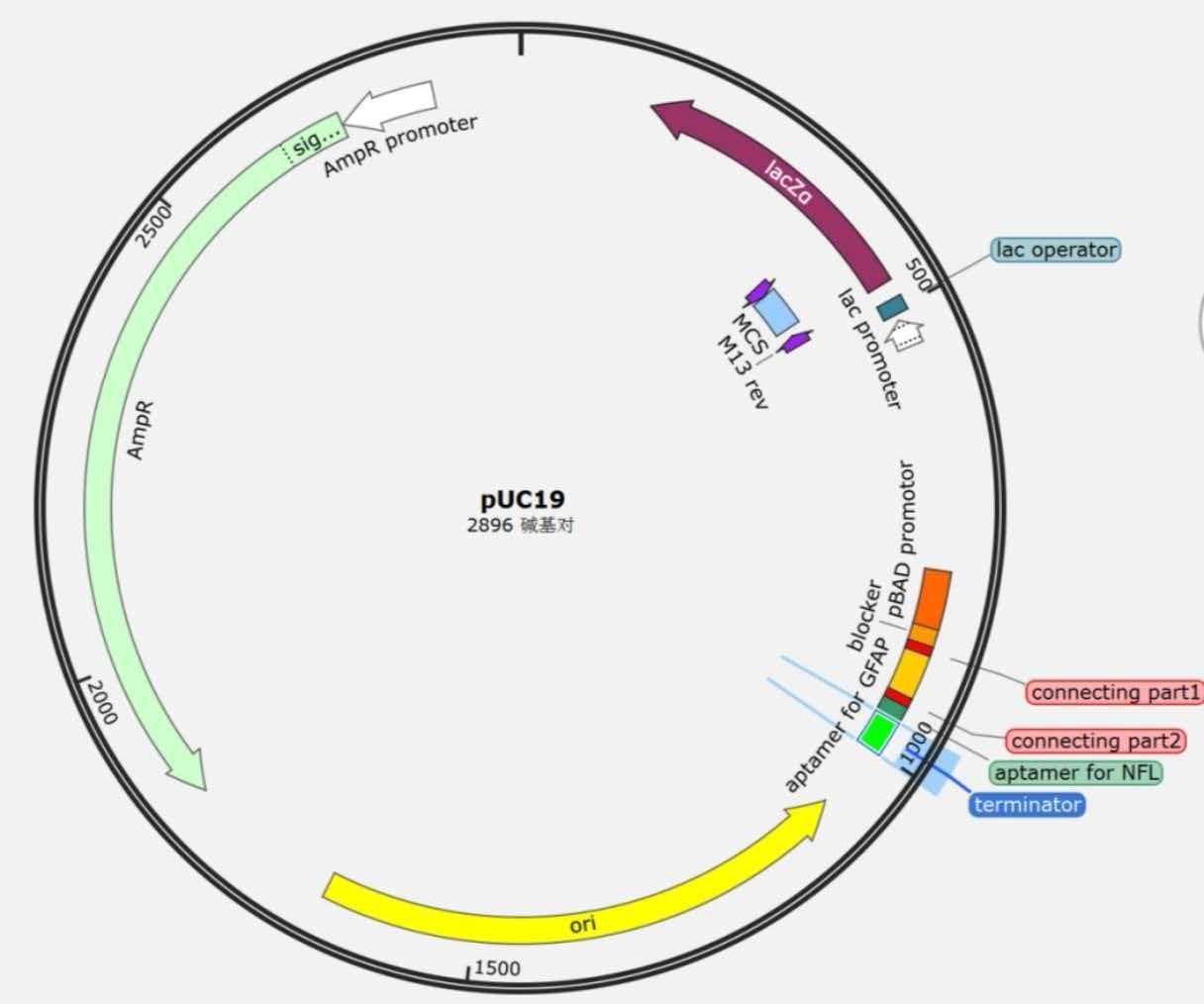
Plasmid 1 (pUC19 backbone) – RNA1 Expression Plasmid
· Resistance: Ampicillin
· Origin of replication: pMB1
· Insert: pBAD promoter (arabinose-inducible) – blocker – (G₄S)₂ linker – GFAP aptamer – (G₄S)₂ linker – NFL aptamer – T7 terminator
· Function: Under arabinose induction, expresses RNA1 (the blocking strand containing dual aptamers)
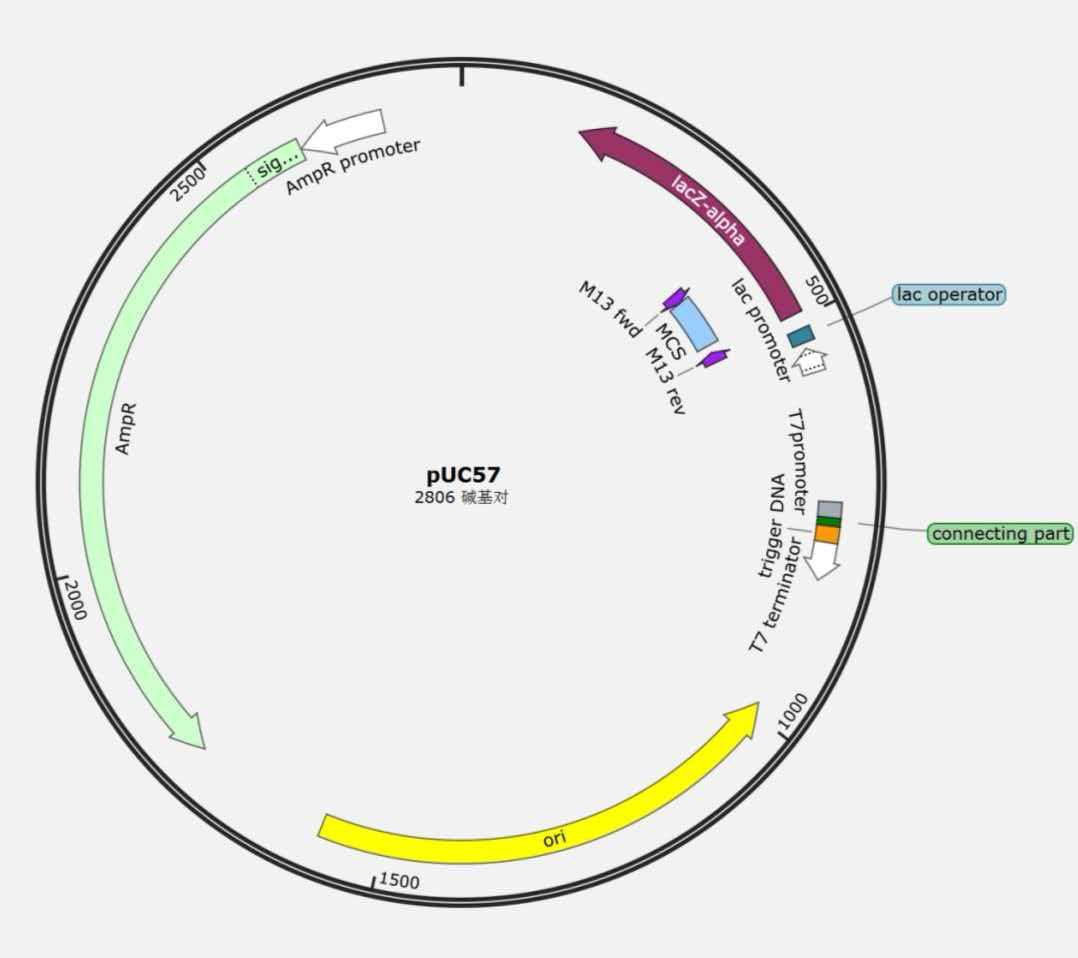
Plasmid 2 (pUC57 backbone) – Reporter Plasmid
· Resistance: Kanamycin
· Origin of replication: pUC
· Insert: T7 promoter – sgRNA – T7 terminator
· Function: Contains the T7 promoter that is blocked by RNA1 in OFF state; upon RNA1 release, transcribes sgRNA
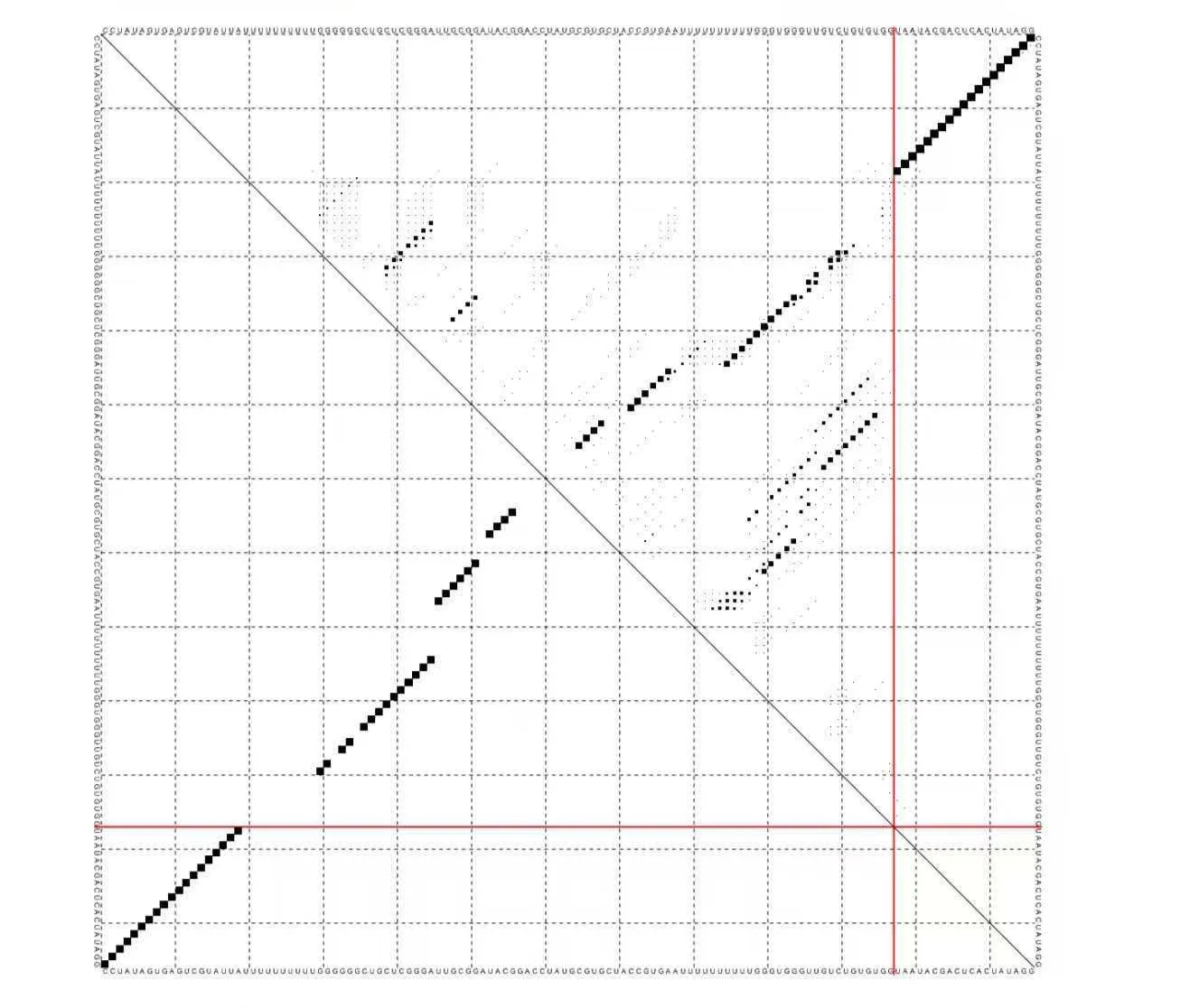
3. Mechanism
OFF state (no target or single target only): RNA1 (containing blocker and dual aptamers) forms a stable duplex with the T7 promoter DNA on the reporter plasmid (ΔG = -27.35 kcal/mol), physically blocking T7 RNA polymerase binding → transcription terminated, no sgRNA produced.
ON state (both NFL and GFAP present):
When NFL and GFAP are both present:
1. NFL binds to the NFL aptamer on RNA1
2. GFAP binds to the GFAP aptamer on RNA1
3. Protein binding induces a conformational change in RNA1, releasing it from the T7 promoter DNA
4. T7 RNA polymerase binds to the promoter and transcribes sgRNA
5. sgRNA can be detected via Cas13-mediated cleavage of a reporter probe, ultimately visualized on a lateral flow test strip.
4. Test Strip Readout Logic (Competitive Format)
| State | sgRNA | Cas13 Activity | Reporter Probe | T Line | C Line | Interpretation |
|---|---|---|---|---|---|---|
| No target / Single target | Absent | Inactive | Intact | Visible | Visible | Negative (no dual target) |
| Dual target present | Present | Active | Cleaved | Invisible | Visible | Positive (NFL+GFAP both present) |
Note: T line: coated with streptavidin to capture biotin-labeled intact reporter probes. C line: coated with anti-FITC antibodies to capture gold-labeled antibodies, serving as a validity control. Invalid result: C line invisible.
5. Key Supporting Data
| Data | Value | Description |
|---|---|---|
| RNA1–T7 binding energy | -27.35 kcal/mol | OFF-state stability |
| NFL docking score | -288.23 | Aptamer–protein binding |
| GFAP docking score | -301.91 | Aptamer–protein binding |
| LOD | 0.57 pM | Sensitivity |
| Linear range | 1–10 nM | Optimal detection range |
| Transcription rate (kₜₓ) | 0.5 nM/s | Key for signal amplification |
6. Detection Workflow
Blood sample + reagents → incubation → Apply to lateral flow test strip → Visual readout
7. References
- Matsumoto M, Ikebukuro K, Tsukakoshi K. Competitive-SELEX discovery of DNA aptamers selective for neurofilament light chain in human plasma. Biochem Biophys Res Commun. 2026;796:153151. (NFL aptamer)
- Cao Y, Lin Y, Chen L, et al. Screening of glial fibrillary acidic protein specific aptamer and application in the development of fluorescent biosensor based on isothermal amplification strategy. Chin J Anal Chem. 2025. (GFAP aptamer)
- Wang X, Shi Z, Qiu Y, Sun D, Zhou H. Peripheral GFAP and NfL as early biomarkers for dementia: longitudinal insights from the UK Biobank. BMC Med. 2024;22(1):192. (NFL+GFAP联合检测依据)
- iGEM Registry. Parts BBa_K5498011, BBa_25ECYPMT, BBa_25T3HMKA.
Background
Alzheimer's disease (AD) affects over 10 million people in China, yet approximately 64% of patients experience delayed diagnosis (ADI 2022). Current diagnostic methods have significant limitations: invasive CSF puncture, costly PET-CT, and subjective cognitive assessments.
AD pathology is heterogeneous, and single biomarkers cannot fully reflect disease status. For example, p-tau217 detection alone may yield false positives due to other diseases or renal dysfunction (Alzheimers Dement 2025; JAMA Neurol 2026). Therefore, synergistic multi-target detection is key to improving diagnostic specificity.
We selected NFL and GFAP as dual targets:
- NFL: reflects axonal injury, elevated in early AD (Matsumoto et al., BBRC 2026)
- GFAP: reflects neuroinflammation, closely linked to AD pathology (Cao et al., Chin. Chem. Lett. 2025)
Our RNA aptamer-based "AND" gate generates a signal only when both targets are present, validated by molecular docking (score -288.23) and RNA structure prediction (ΔG = -27.35 kcal/mol).
Core Highlights
Completed Design Highlights
- 🎯 Dual-target synergy: Simultaneous detection of NFL (axonal injury) and GFAP (neuroinflammation)
- 🔗 "AND" gate logic: Signal generated only when both targets are present, eliminating single-target false positives
- 🔒 OFF-state stability: RNA1 binds T7 promoter with ΔG = -27.35 kcal/mol (RNAcofold)
- 🧪 Aptamer affinities: NFL Kd = 8.1 nM / GFAP Kd = 0.62 μM (literature)
- 🔬 Molecular docking: HDOCK scores NFL -288.23 / GFAP -301.91
- 🧬 Full-length DNA sequence: Designed and available in Wiki
- 📊 Predicted performance: LOD 0.57 pM, linear range 1–10 nM (ODE modeling)
- ⚙️ Parameter analysis: kₜₓ and γ identified as key levers for signal amplification
Product & Translation Highlights
- 🏠 Lateral flow test strip: Simple, home-based operation
- 👁️ Visual readout: Results in 5–15 min, no equipment needed
- 💰 Low cost: Projected <5 RMB per test at scale
- 💬 User-aligned: Meets "like a pregnancy test" expectation from interviews
- 🧩 Modular design: Aptamers replaceable for other targets
- 🌍 Social impact: Addressing delayed AD diagnosis with accessible screening
Project Progress
| Phase | Status | Source |
|---|---|---|
| Literature review & target selection | ✅ Completed | Wang et al., BMC Med 2024 |
| NFL aptamer sequences obtained | ✅ Completed | Matsumoto et al., BBRC 2026 |
| GFAP aptamer sequences confirmed | ✅ Completed | Cao et al., Chin J Anal Chem 2025 |
| Full-length DNA sequence designed | ✅ Completed | Team design (sequence in Wiki) |
| OFF-state RNA structure prediction | ✅ Completed | RNAcofold, ΔG = -27.35 kcal/mol |
| Molecular docking validation | ✅ Completed | HDOCK: NFL -288.23 / GFAP -301.91 |
| Kinetic modeling & "AND" gate validation | ✅ Completed | ODE model, four-curve comparison |
| Detection performance analysis | ✅ Completed | LOD = 0.57 pM, linear range 1–10 nM |
| Parameter sensitivity analysis | ✅ Completed | k_off ≤ 10⁻⁵ sufficient; k_tx and γ are key levers |
| User questionnaire survey | ✅ Completed | Online survey (N=93), March 2026 |
| Experimental validation | ⏳ Pending lab access | DNA synthesis & in vitro validation required |
| Prototype development | ⏳ Planned | Further engineering optimization needed |
Future Plans
Based on team discussions, the final product format has been determined: lateral flow test strip with colloidal gold visualization, enabling home-based point-of-care testing (POCT).
Test Strip Structure
- Sample pad: Accepts the test sample (fingerstick blood)
- Conjugate pad: Contains colloidal gold-labeled detection molecules (aptamers or reporters)
- Nitrocellulose membrane (NC membrane): Coated with test line (T line) and control line (C line)
- Absorbent pad: Provides capillary driving force
Workflow
- Sample collection: Fingerstick blood (10–50 μL)
- One-step application: Apply blood directly to the sample pad
- Lateral flow: Liquid migrates by capillary action, reacting with colloidal gold conjugates
- Result readout: Visual inspection after 5–15 min
Positive: Both T line and C line appear (purple-red) → NFL and GFAP both present
Negative: Only C line appears → no target or single target only
Invalid: C line absent
Product Advantages
- ✅ No equipment needed: Visual readout, suitable for home use
- ✅ Simple operation: One-step application, no training required
- ✅ Rapid: Results in 5–15 min
- ✅ Low cost: Projected <5 RMB per test at scale
- ✅ Aptamer compatible: Nucleic acid aptamers can be stably conjugated to gold nanoparticles
- ✅ Fulfills user expectations: Achieves the "like a pregnancy test" experience
Development Roadmap
- Short-term: Optimize aptamer-gold conjugation conditions (pH, concentration)
- Mid-term: Assemble complete test strips, test sensitivity and specificity
- Long-term: Develop lyophilized reagents for room-temperature storage and transport
