Experimental Design
Engineering and validation pipeline of CradleShield
Experimental Design
From plasmids to functional validation
🧬
1. Plasmid Construction & Validation
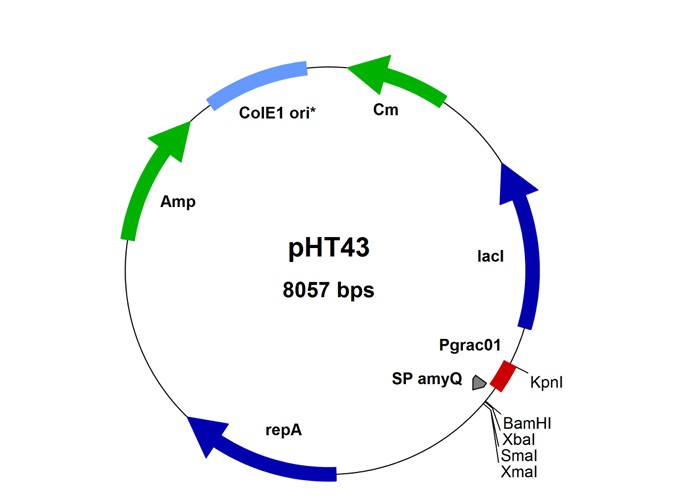
Goal: Construct three plasmids (pHT43‑butyrate, pAX01‑butyrate‑dal, pK18mobsacB‑Δdal) in E. coli DH5α and verify sequences.
🔧 Assembly
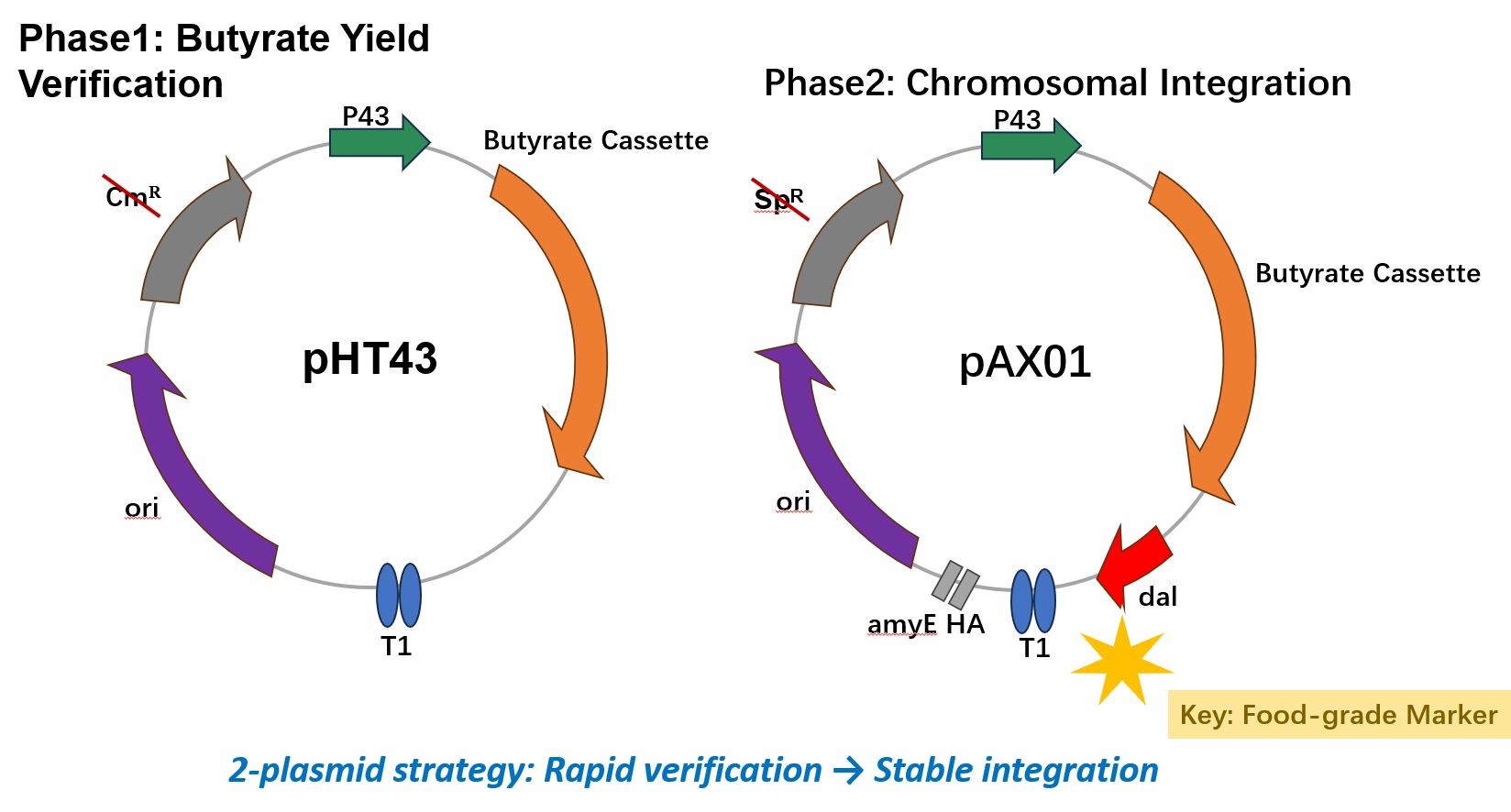
- Golden Gate: pHT43‑butyrate (P43 + RBS + butyrate cluster + terminator).
- Gibson Assembly: pAX01‑butyrate‑dal and pK18mobsacB‑Δdal.
- Transformation into E. coli DH5α with antibiotic selection (chloramphenicol, spectinomycin, kanamycin).
🔍 Validation
- Colony PCR, restriction digestion, and Sanger sequencing.
- Pass rate: ≥95% sequence identity, ≥5 clones per plasmid stored.
🧫
2. Δdal Auxotrophic Strain Construction
Goal: Knock out the dal gene in B. subtilis 168 to create a D‑alanine auxotroph.
⚡ Electroporation & Screening
- Competent cells prepared with glycine; electroporation of linearized pK18mobsacB‑Δdal.
- Single‑crossover integrants selected on kanamycin plates.
- Double‑crossover (Δdal) selected on sucrose plates (loss of kanamycin resistance).
✅ Validation
- PCR: no dal band.
- Auxotrophy test: growth only on M9 + D‑alanine.

🧪🧬🔬
🧩
3. Chromosomal Integration of Butyrate Cluster
Goal: Integrate the butyrate synthesis cassette into the amyE locus of Δdal strain.
- Electroporation of pAX01‑butyrate‑dal into Δdal competent cells.
- Selection on LB + spectinomycin + D‑alanine.
- Marker loss: passaging without antibiotics, then screening for spectinomycin sensitivity.
- PCR verification of integration at amyE and dal complementation test.
🌱
4. Spore Induction & Purification
- Culture in SP medium with Mn²⁺ for 48 h (sporulation rate ≥90%).
- Purification: ethanol treatment (75%, 30 min) kills vegetative cells.
- Wash and resuspend in saline → purified spore preparation.
- Antibiotic resistance: Spores grow on LB + cephalosporins/amoxicillin.
- Stability: ≥90% germination after 7 days at 25°C.
🧪🌡️❄️
📊
5. Functional Validation
🧪 Biochemical
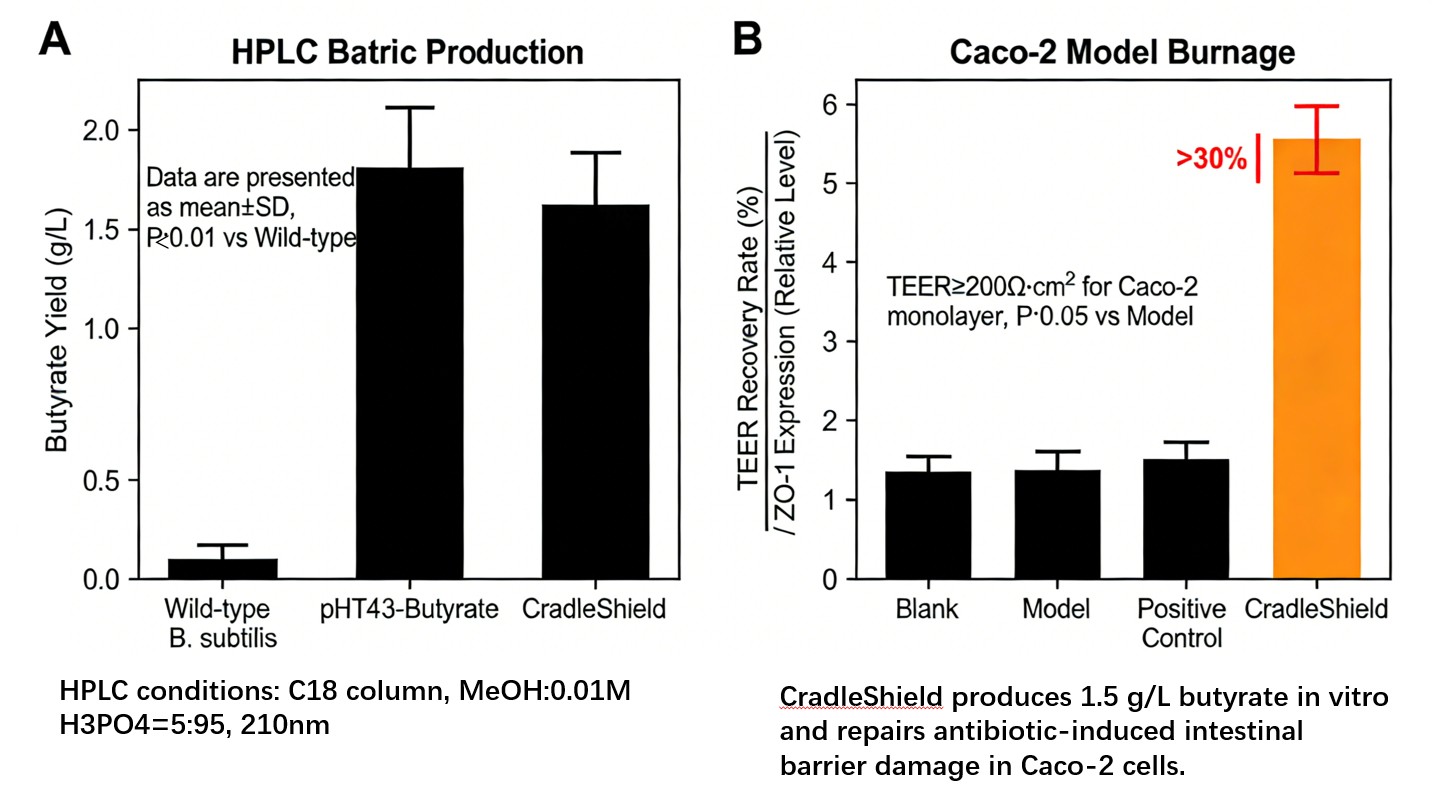
HPLC quantification of butyrate: ≥1.5 g/L (wild‑type <0.1 g/L).
🧫 Cellular (Caco‑2)
- TEER increase ≥30%
- FITC‑dextran permeability decrease ≥40%
🧬 Molecular
HDAC8 inhibition ≥50%; Slc26a3, ZO‑1, Occludin up‑regulated ≥2‑fold.
🐭 In vivo (mouse AAD model)
- Diarrhea index decreased by 40‑60%.
- Colon histology: reduced mucosal damage.
- Tight junction proteins (ZO‑1/Occludin) restored.
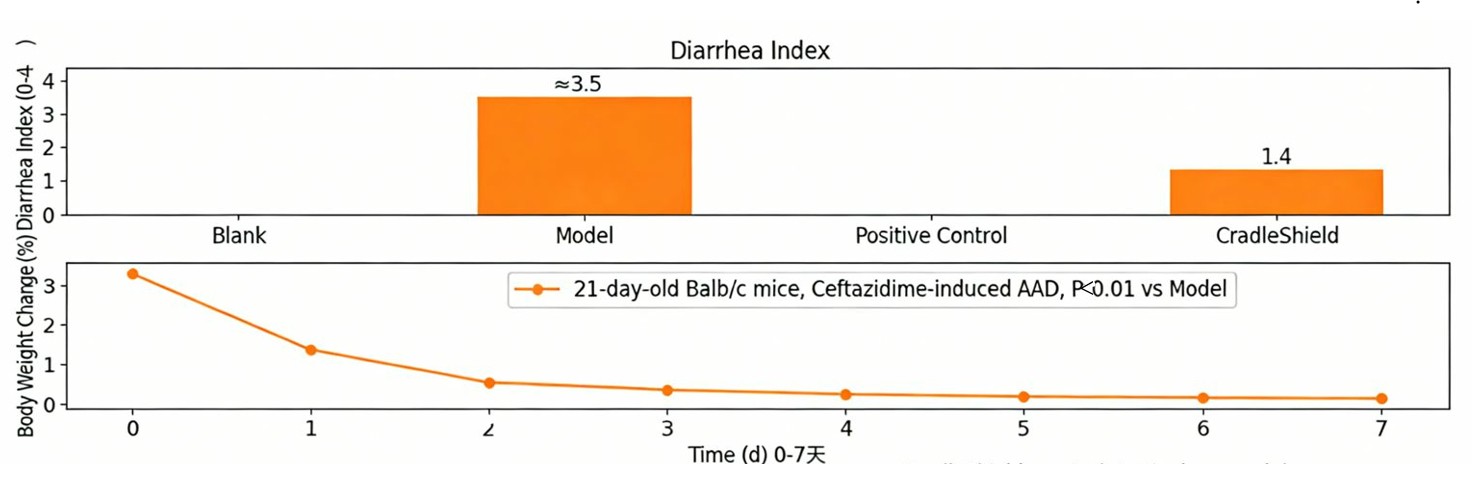
🐭📈🧫
🛡️
6. Biosafety Validation
🌍 Environmental self‑limitation
No growth on M9 (without D‑alanine) or in water/soil extracts.
🧬 Horizontal gene transfer
No transconjugants/transformants detected with E. coli or pathogens.
🐁 Acute toxicity
Mice gavaged with 10¹⁰ CFU for 7 days showed no abnormal symptoms, normal weight gain, and no organ pathology.
🔬🧪🧫🛡️