Genetic Circuit and Parts
Design and construction of plasmids and genetic elements
Genetic Circuit & Parts
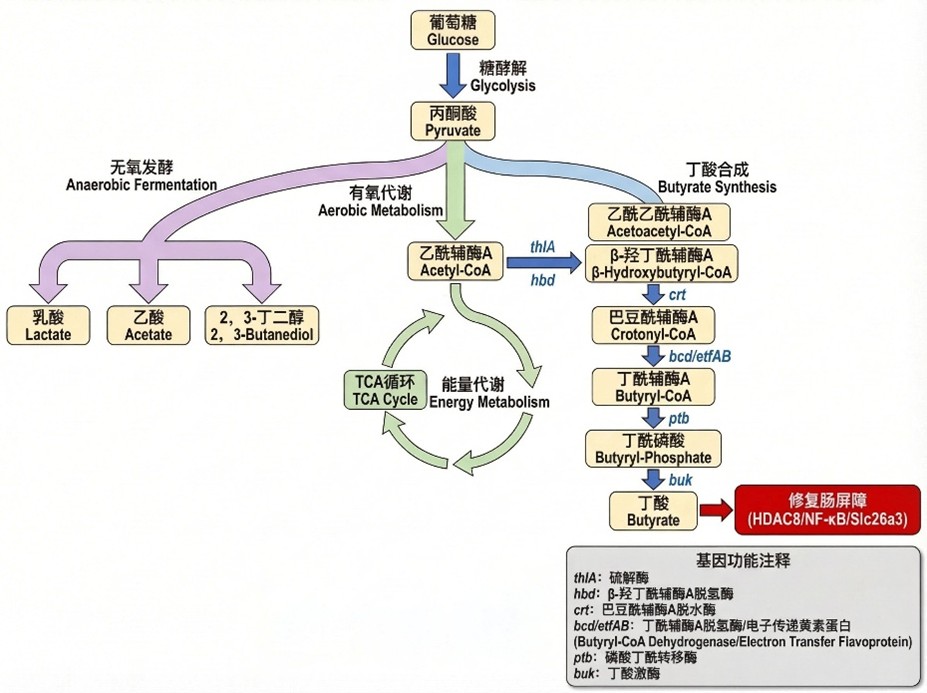
Modular design of the butyrate synthesis pathway
Design Concept
⚠️ Clinical Problem
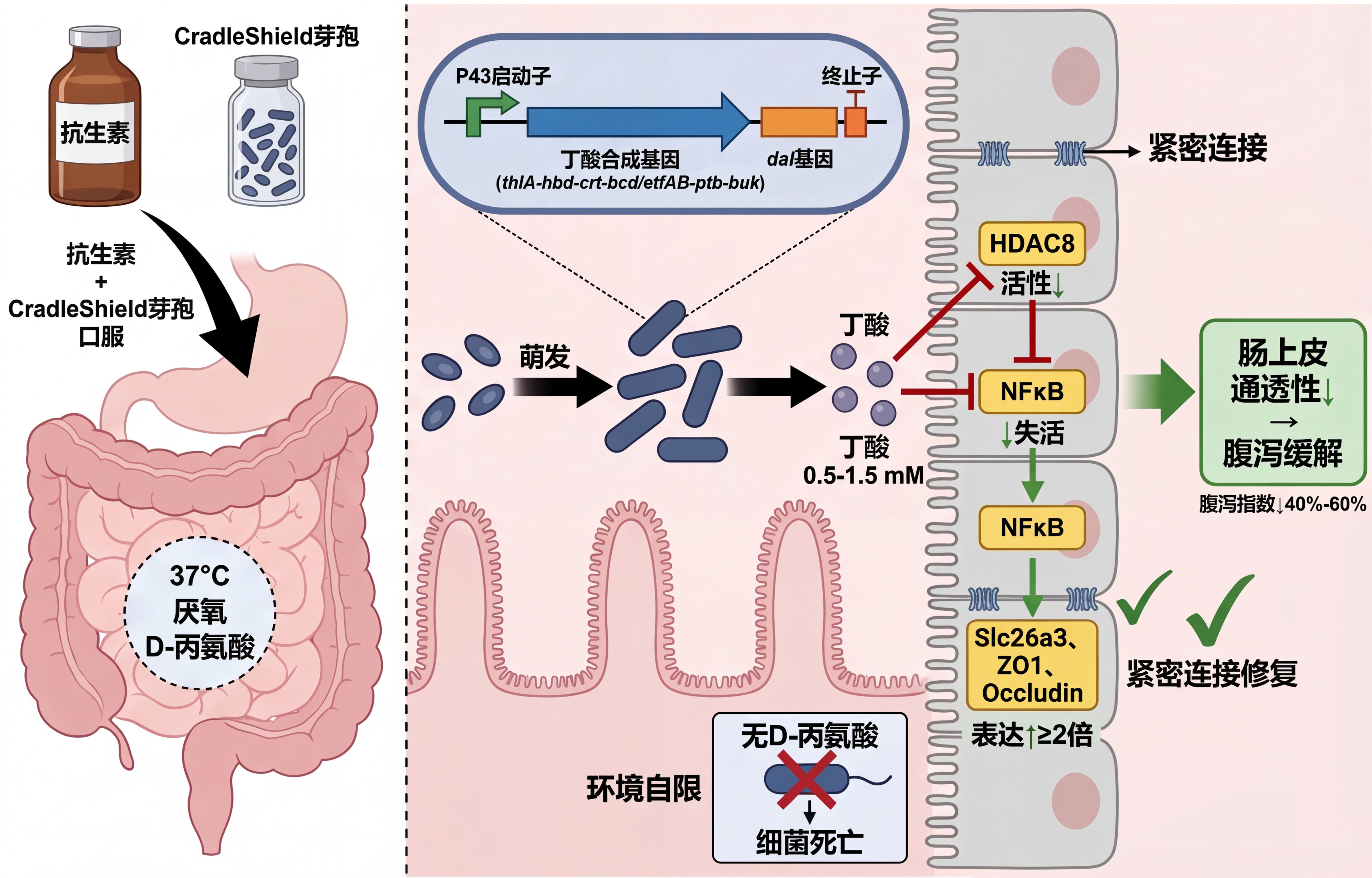
Antibiotic‑associated diarrhea (AAD) in children is caused by the destruction of butyrate‑producing gut microbiota, leading to impaired tight junctions. Current probiotics are antibiotic‑sensitive, require cold chain, and act too slowly.
🦠 Chassis & Functional Molecule
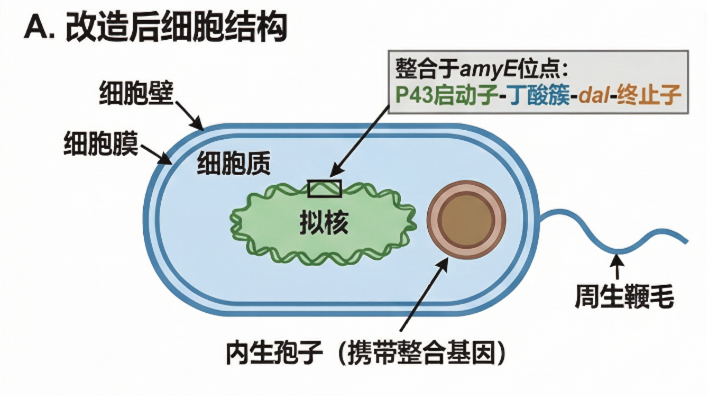
- Chassis: Bacillus subtilis 168 spores – naturally resistant to cephalosporins/amoxicillin, room‑temperature stable, no gut colonization.
- Molecule: Butyrate (0.5–1.5 mM) – repairs tight junctions via HDAC8/Slc26a3 pathway, no colonization lag.
- Safety: dal auxotrophy – D‑alanine dependent growth, environmental self‑limitation.
🧠 Logic Layer: Sense–Compute–Respond
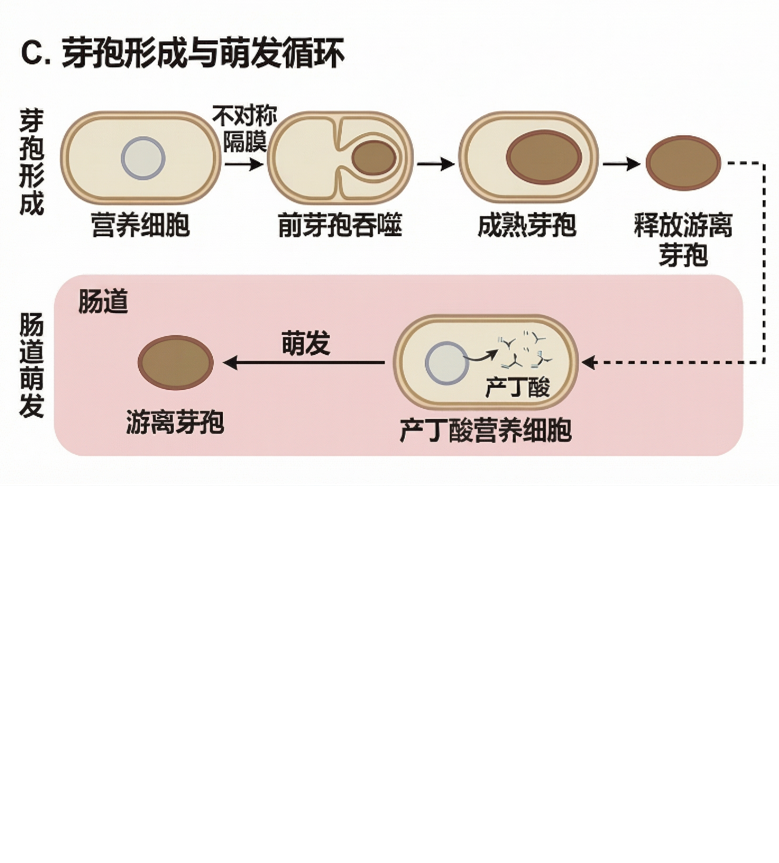
- Sense: Spore germination in the gut (37°C, anaerobic, D‑alanine present) – no external inducer.
- Compute: Constitutive strong promoter P43 drives expression – simple, stable.
- Respond: Dual output – butyrate synthesis and dal complementation (food‑grade selection & self‑limitation).
📐 General Design Rules
🧬 GenBank format with full annotations.
✂️ BsaI sites removed internally for Golden Gate.
🧪 Codon‑optimized butyrate cluster (GC% 45–55).
🔗 Spacing: P43‑RBS 6 bp, RBS‑ATG 4 bp, terminator after TAA.
🧫 Shuttle plasmids for E. coli and B. subtilis.
🧩 Key Plasmids
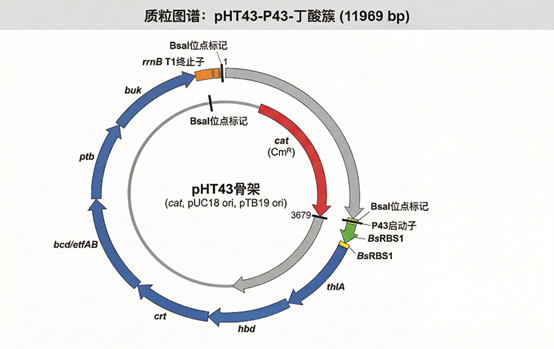
🧬 pHT43‑butyrate (Phase 1 validation)
- Backbone: pHT43 shuttle vector, Cmᴿ.
- Expression cassette: P43‑RBS1‑butyrate cluster‑rrnB T1.
- Size: 11,969 bp.
- Assembly: Golden Gate (BsaI).
- Genes: thlA, hbd, crt, bcd/etfAB, ptb, buk with Gly‑Ser linkers.
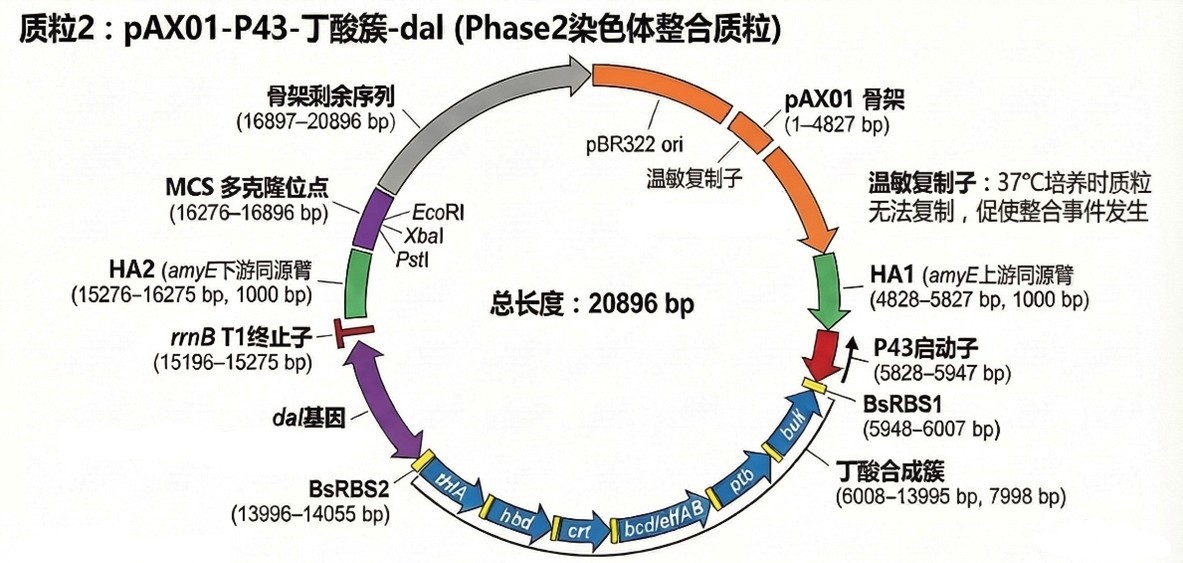
🧩 pAX01‑butyrate‑dal (Phase 2 integration)
- Backbone: pAX01 integration vector, Spcᴿ (lost after integration).
- Cassette: amyE‑HA1 – P43‑RBS1‑butyrate‑RBS2‑dal‑terminator – amyE‑HA2.
- Size: 20,896 bp.
- Assembly: Gibson Assembly.
- Integration locus: amyE (chromosomal).
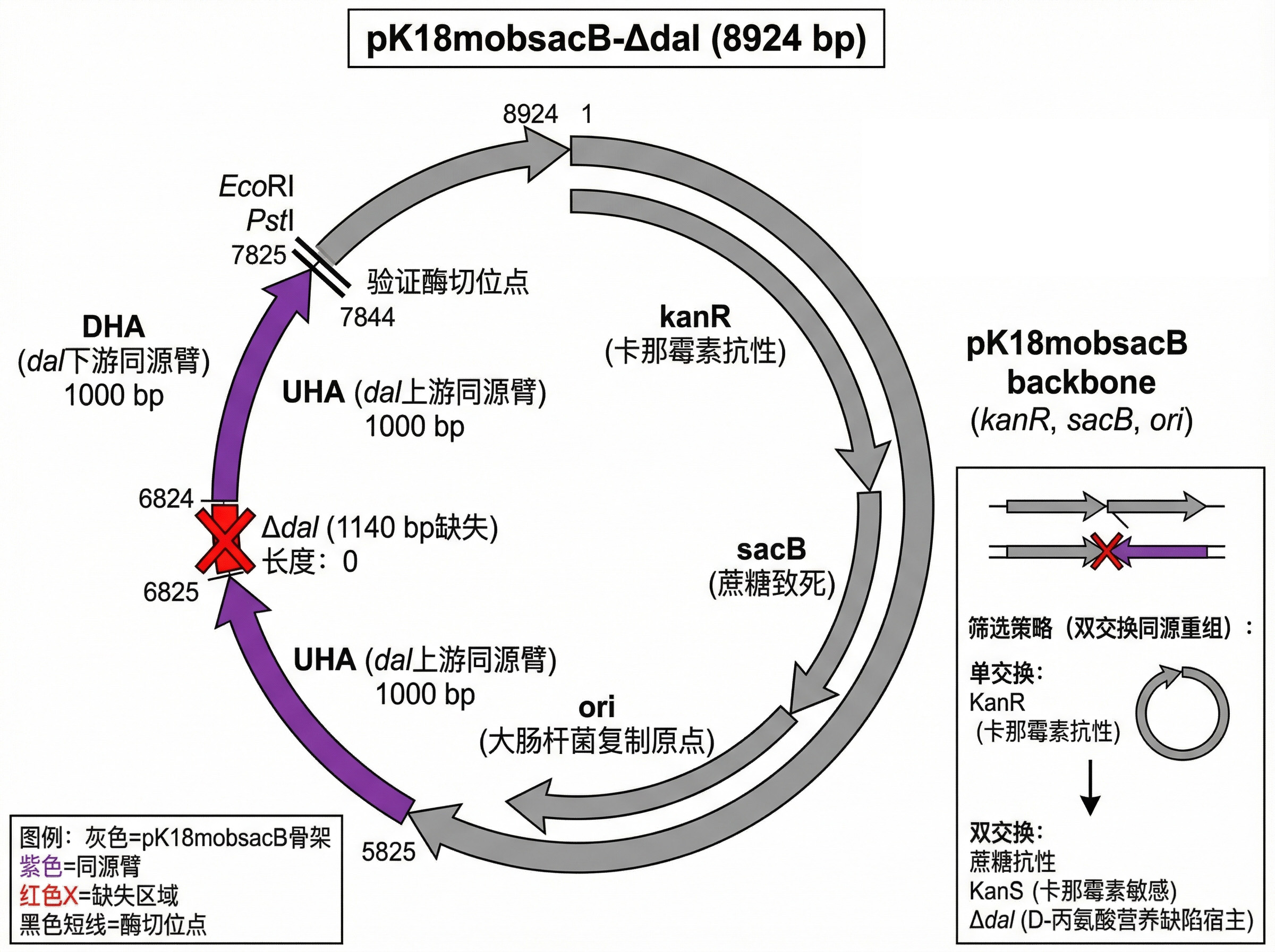
✂️ pK18mobsacB‑Δdal (Gene knockout)
- Backbone: pK18mobsacB, Kanᴿ, sacB counter‑selection.
- Knockout cassette: dal upstream‑Δdal‑dal downstream.
- Size: 8,924 bp.
- Assembly: Gibson Assembly.
- Function: Creates Δdal auxotrophic strain.
🔄 Engineering Iteration (DBTL)
🧪 Phase 1 (Plasmid)
Design: pHT43‑butyrate, high‑copy.
Build: Golden Gate assembly.
Test: HPLC butyrate ≥1.5 g/L.
Learn: Plasmid unstable, needs antibiotics → move to integration.
🔧 Phase 2 (Chromosomal)
Design: pAX01‑butyrate‑dal with amyE arms.
Build: Gibson Assembly, transform into Δdal.
Test: PCR verify integration, butyrate ≥1.2 g/L.
Learn: Stable, no antibiotic markers → final strain.
⚙️ Physical Construction Steps
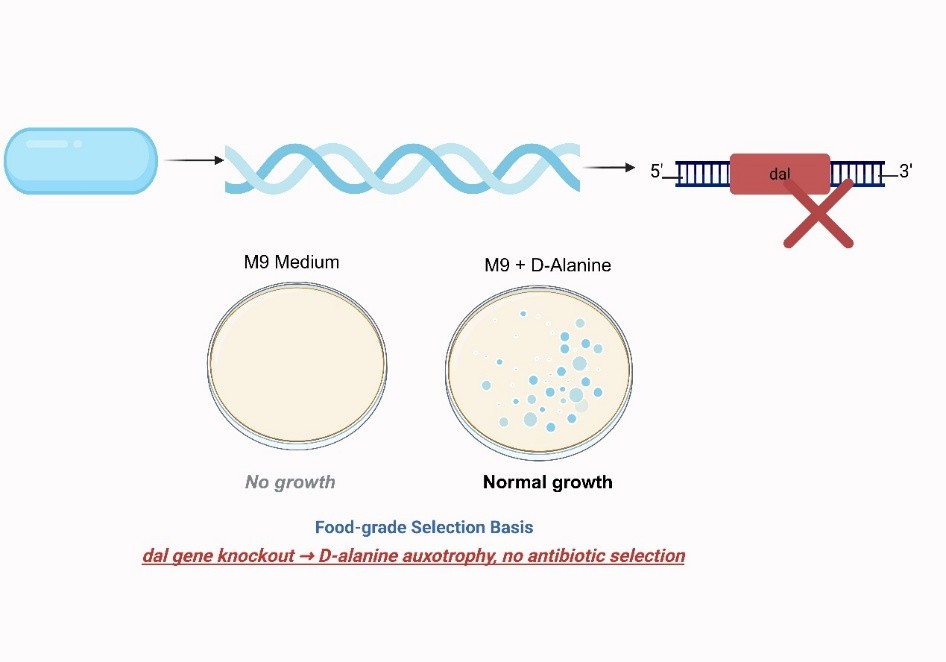
Δdal Host Construction
Knockout of dal via pK18mobsacB‑Δdal → D‑alanine auxotrophy.
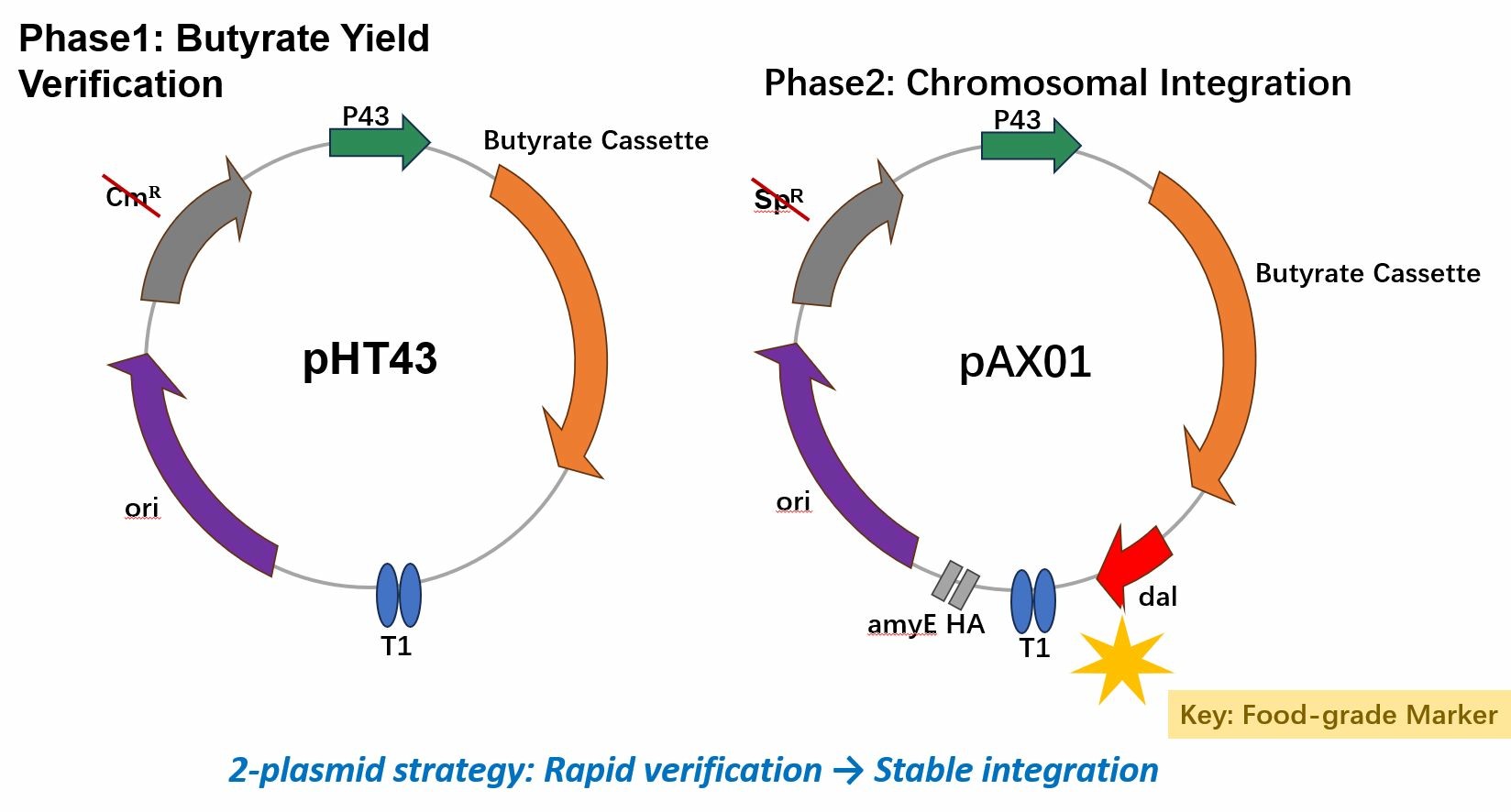
Plasmid Construction
pHT43‑butyrate (Phase 1) and pAX01‑butyrate‑dal (Phase 2).
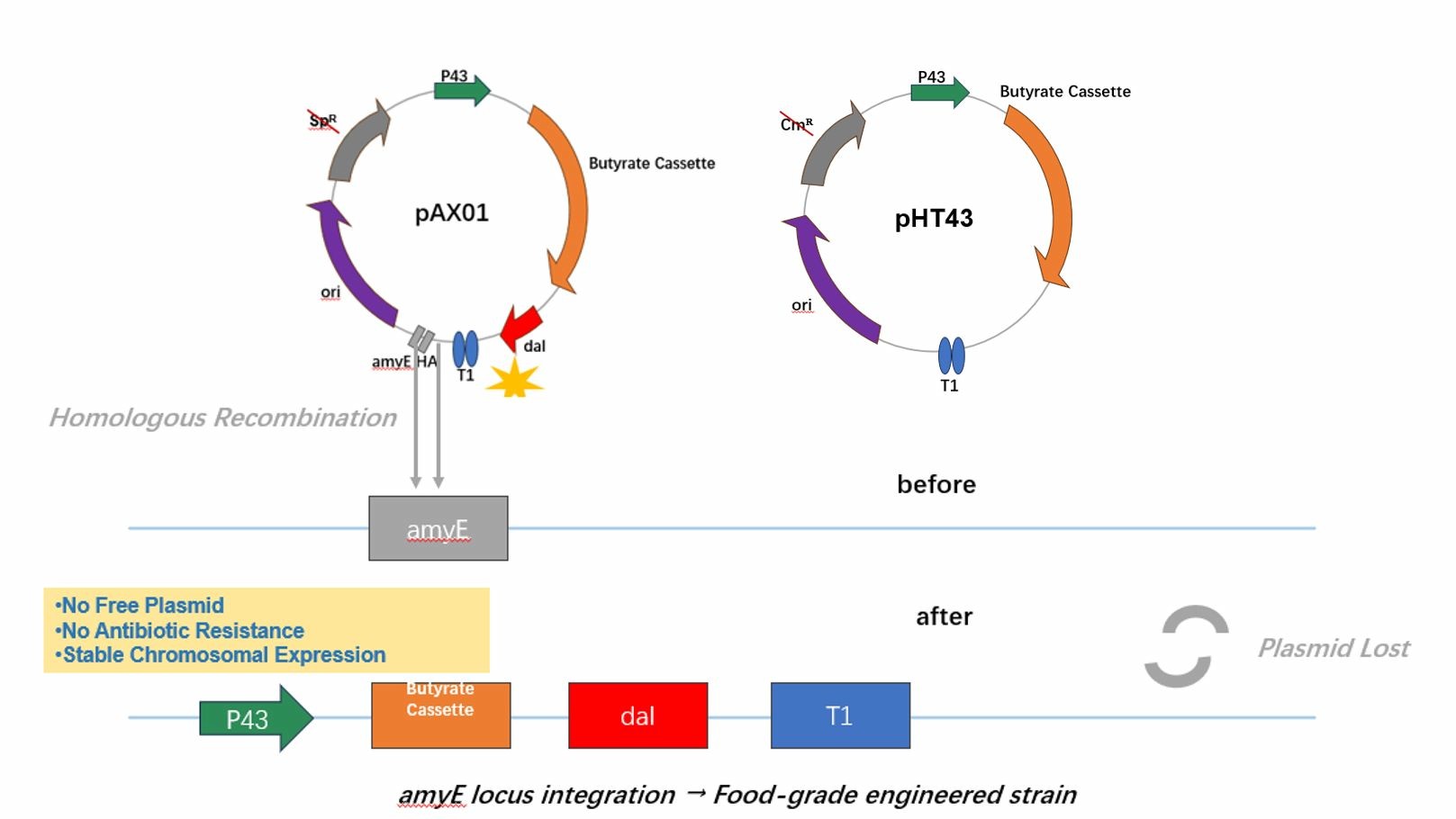
Chromosomal Integration
Integration into amyE → CradleShield (Δdal::amyE‑P43‑butyrate‑dal).