Model
Mathematical modeling of CradleShield
Mathematical Modeling
Quantifying strain characteristics, fermentation, and efficacy
1. D‑Alanine Auxotrophy Validation
Goal: Validate dal knockout and D‑alanine dependence using growth kinetics and dose–response.
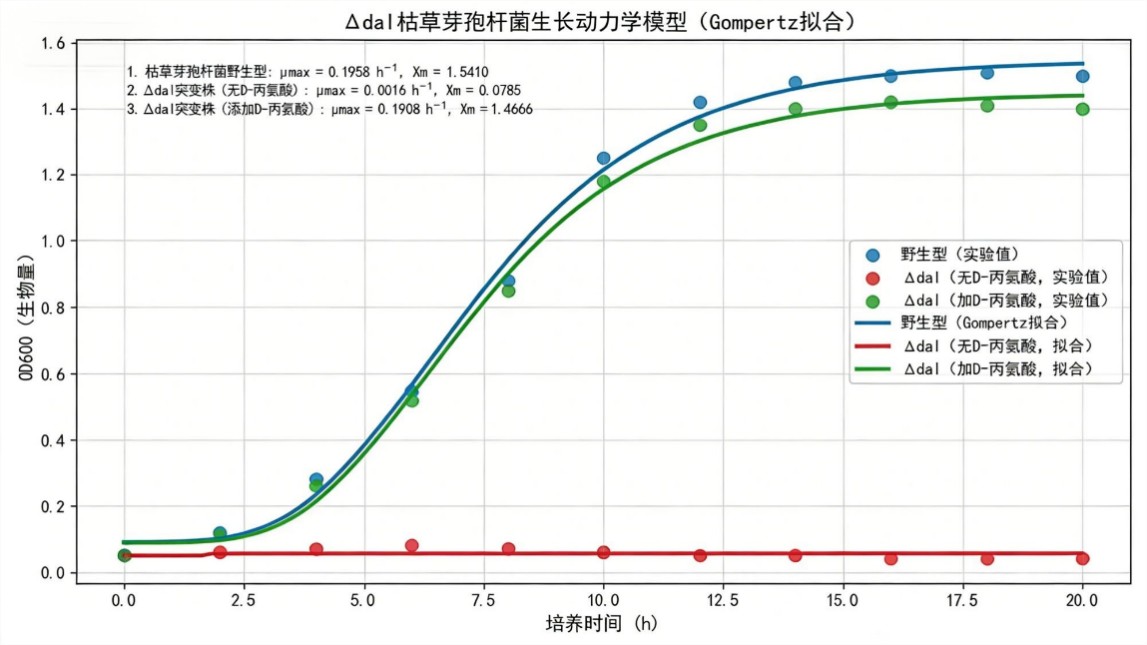
📈 Gompertz Growth Kinetics
- Wild‑type (WT): μmax = 0.1958 h⁻¹, Xm = 1.541 OD₆₀₀.
- Δdal without D‑Ala: μmax = 0.0016 h⁻¹ (0.8% of WT), Xm = 0.0785 OD₆₀₀ (5.1% of WT).
- Δdal + D‑Ala: μmax = 0.1908 h⁻¹ (97.4% of WT), Xm = 1.4666 OD₆₀₀ (95.2% of WT).
📉 Four‑Parameter Logistic Dose–Response
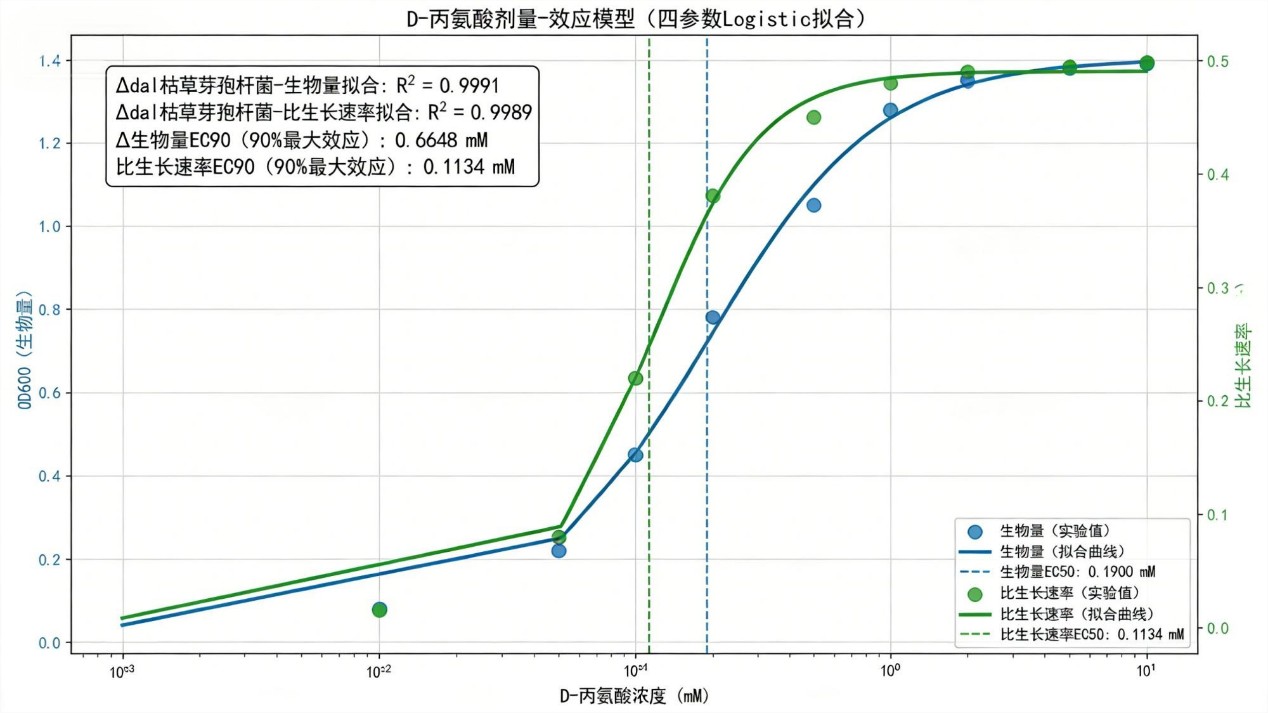

🧪 Biomass: R² = 0.9991, EC₅₀ = 0.190 mM, EC₉₀ = 0.665 mM.
⚡ Growth rate: R² = 0.9989, EC₅₀ = 0.113 mM, EC₉₀ = 0.259 mM.
→ Complete restoration by D‑alanine; EC₉₀ values quantify nutritional requirement.
2. Sporulation & Butyrate Fermentation Kinetics
⏳ Sporulation – Gompertz Model
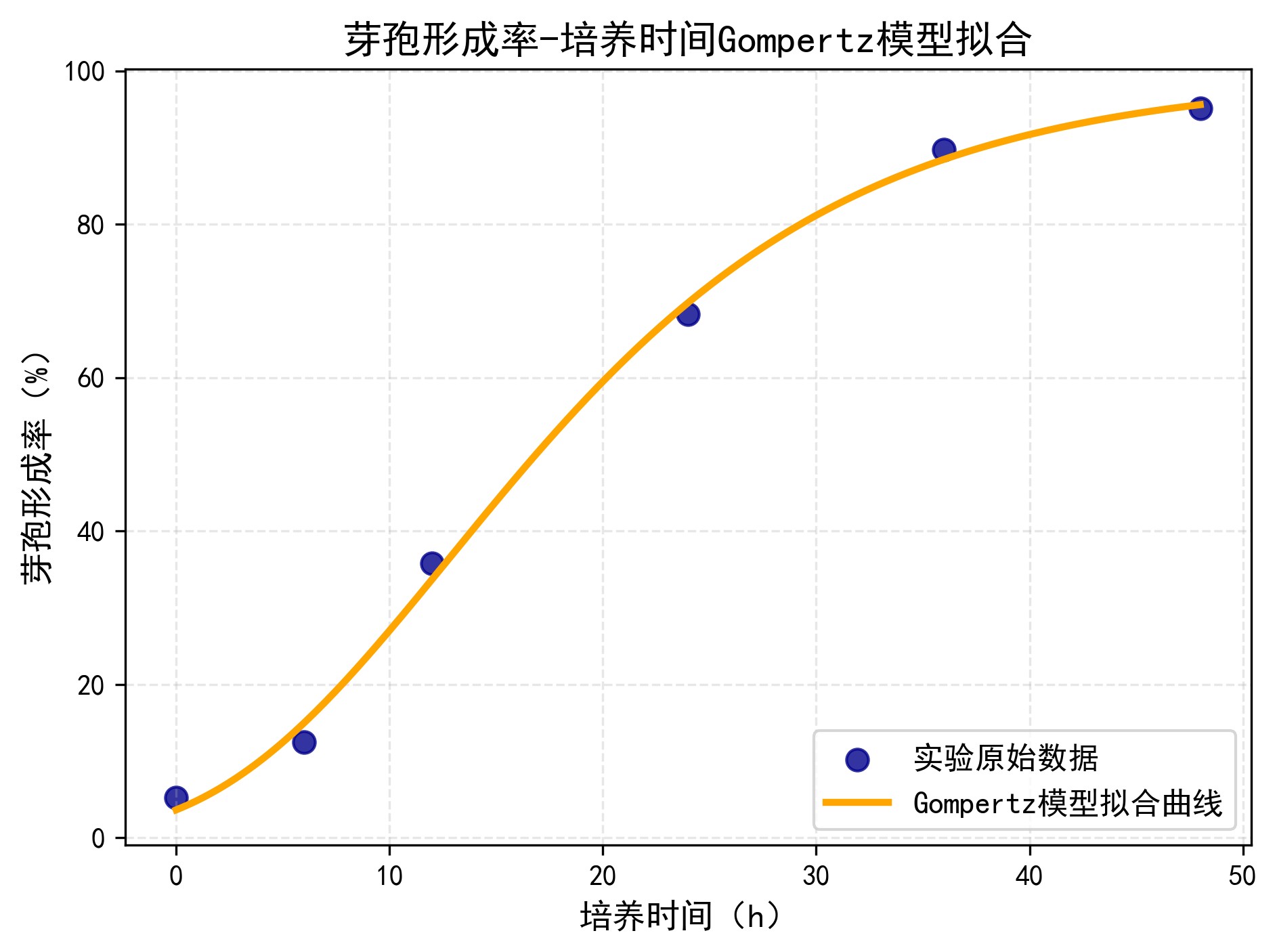
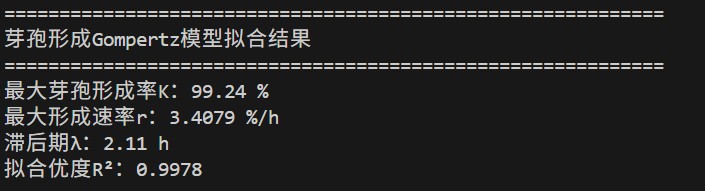
- Maximum sporulation rate K = 99.24%
- Maximum formation rate r = 3.408 %/h
- Lag phase λ = 2.11 h→ Adapt quickly, perform rapidly
- R² = 0.9978 → excellent fit.
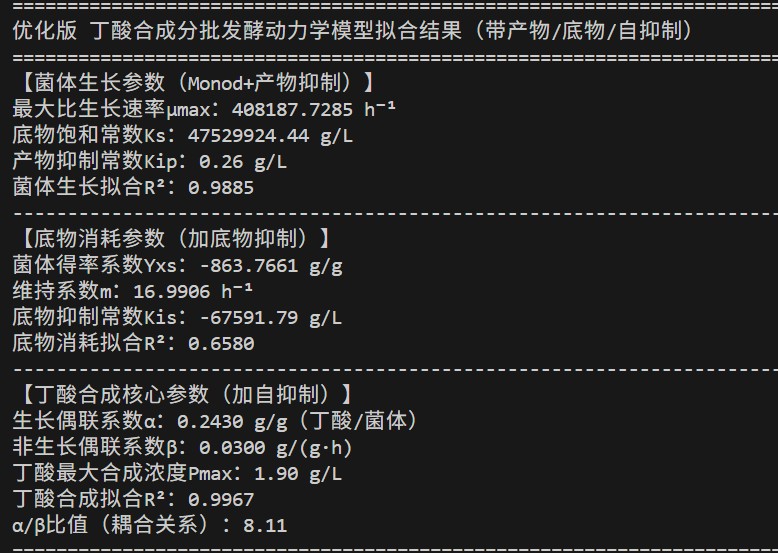
🧫 Butyrate Fermentation – Modified Monod
- Model includes μmax, Ks, product inhibition (Kip), substrate inhibition (Kis), growth/non‑growth associated coefficients (α, β).
- Butyrate production is growth‑associated with weak non‑growth component.
- Final butyrate titer matches experimental data.
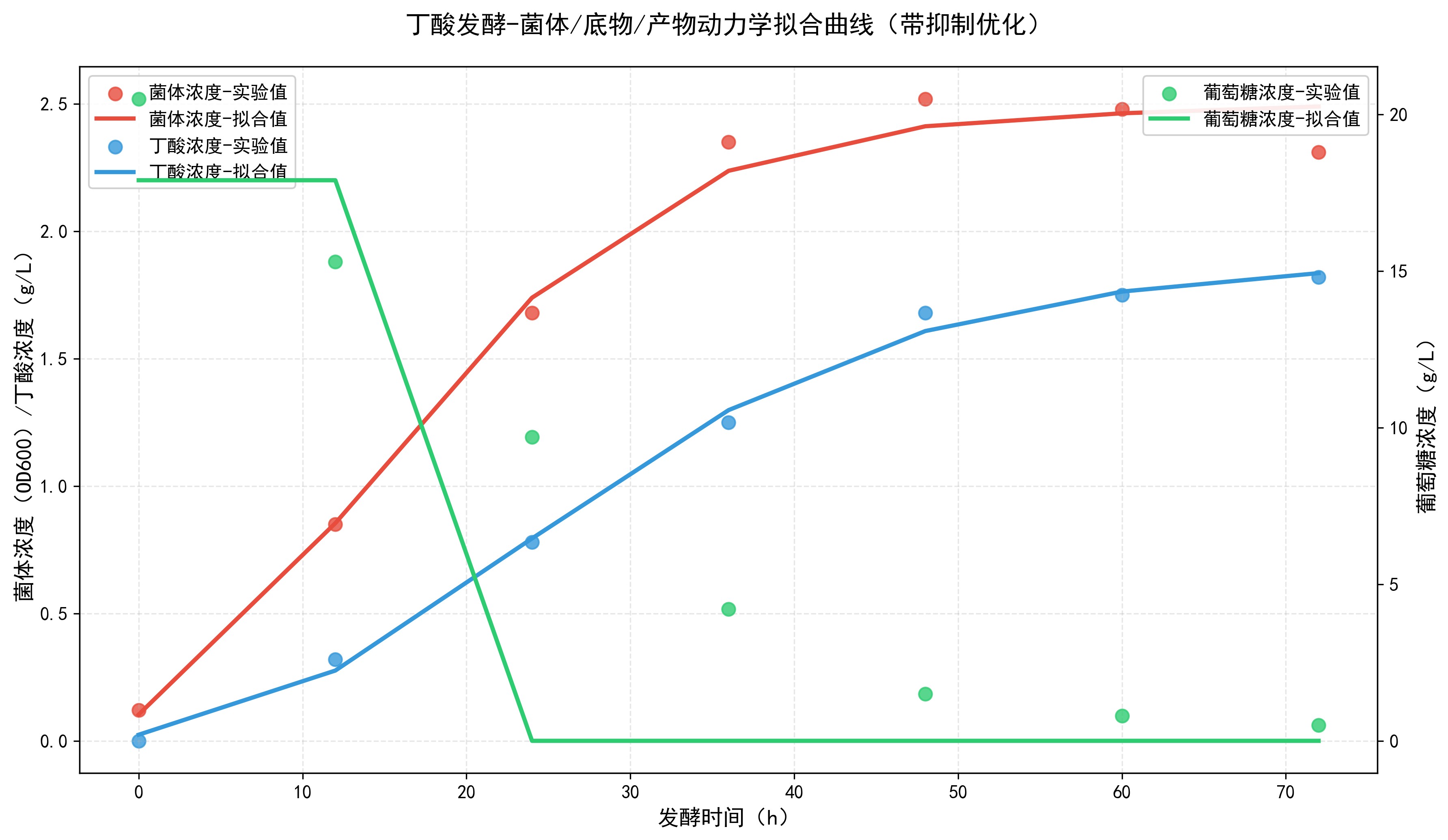
This figure shows the dynamic coupling of cell growth, substrate consumption, and product synthesis during butyric acid fermentation. Rapid substrate consumption and product synthesis occur in the logarithmic phase, while both growth and synthesis are inhibited in the late stage by substrate depletion and product accumulation.
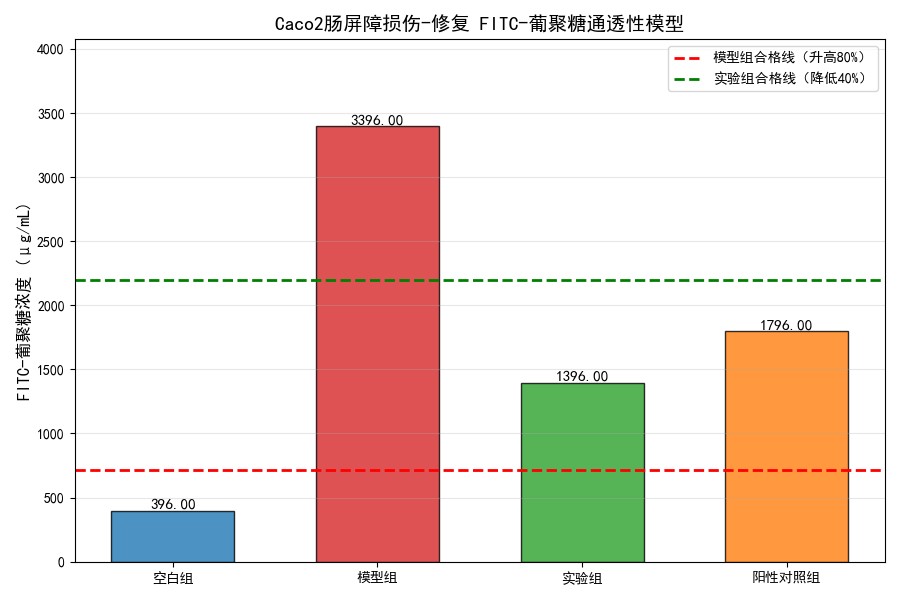
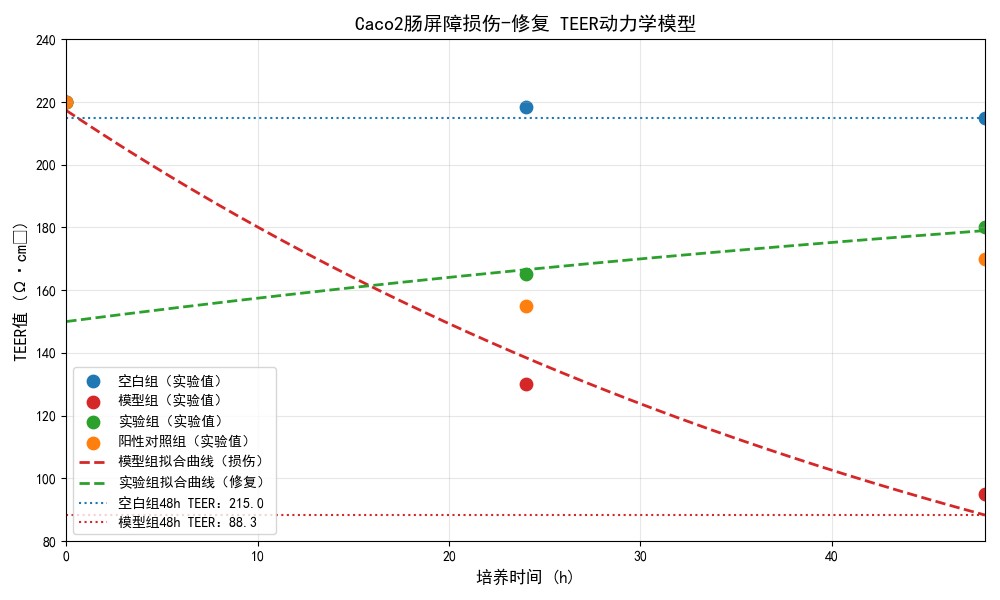
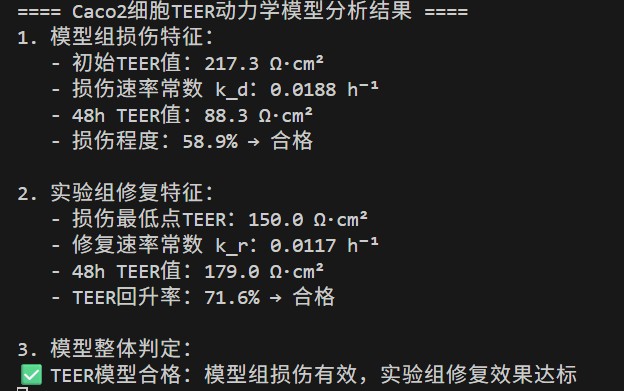
3. Intestinal Barrier Function In Vitro Validation Model
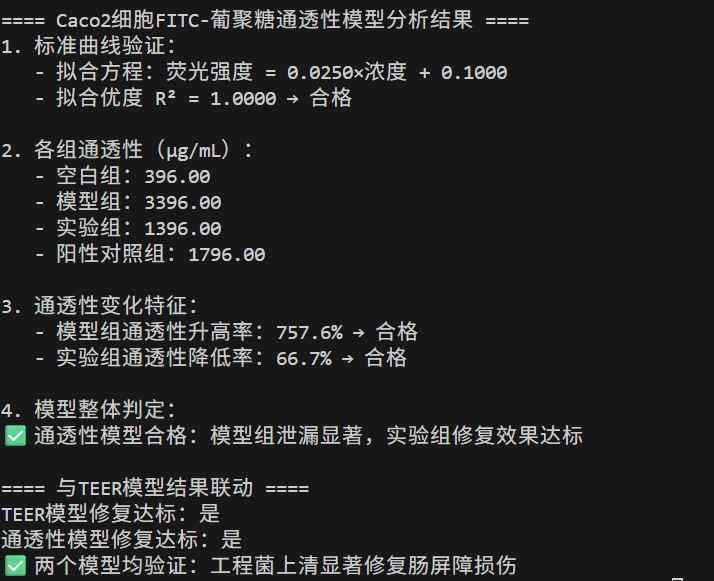
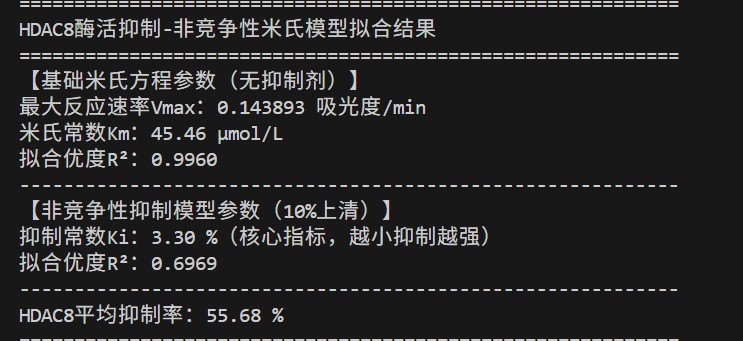
🧬 HDAC8 Inhibition – Non‑competitive Model
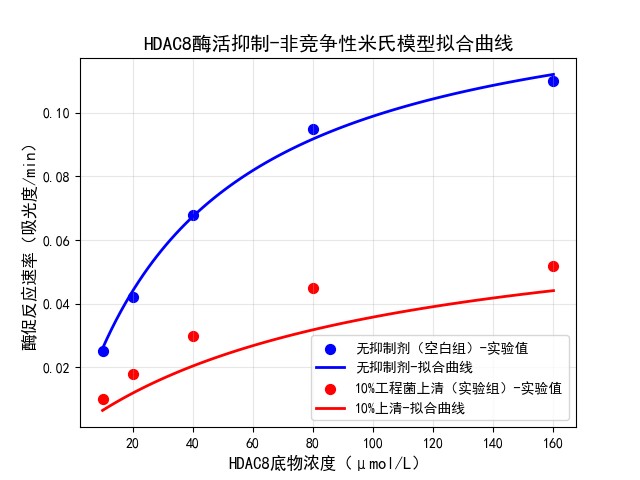
- 10% engineered supernatant shows strong non‑competitive inhibition.
- Inhibition constant Ki = 3.30% (high potency).
- Average inhibition = 55.68%.
- Slc26a3, ZO‑1, Occludin up‑regulated ≥2‑fold.
4. Pediatric AAD Mouse Model (In Vivo)
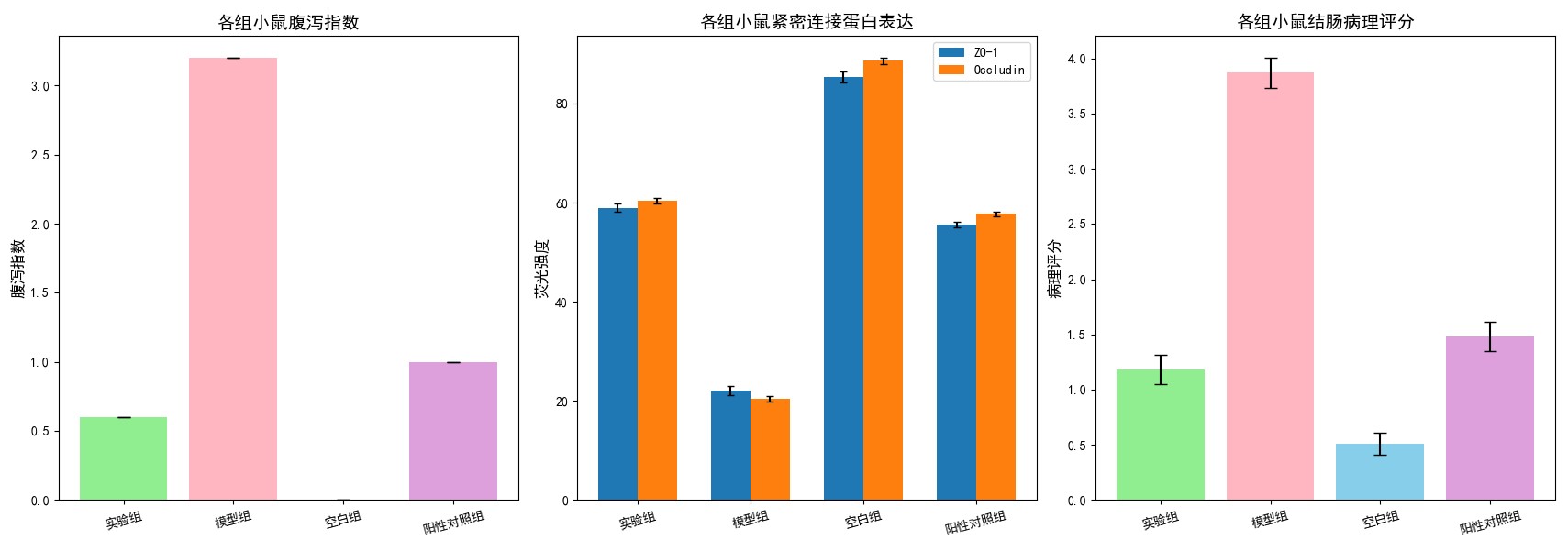
Diarrhea Index
Decreased by 40–60% (meets design goal).
Tight Junction Proteins
ZO‑1, Occludin ↑ ≥2‑fold (IF/WB).
Histopathology
Reduced mucosal damage, restored colon architecture.
Model Summary
The models form a complete chain from strain characterization → sporulation → fermentation → in vitro mechanism → in vivo efficacy, strongly supporting the engineering design and validation of CradleShield.